Glycosaminoglycan Protein Interactions: Molecular Modeling and Simulation Methods
|
|
|
- Isabella Booker
- 6 years ago
- Views:
Transcription
1 Glycosaminoglycan Protein Interactions: Molecular Modeling and Simulation Methods Philip D. Mosier Senior Scientist and Co Director, SPG PEG Computational Chemistry and Biology Core Department of Medicinal Chemistry Virginia Commonwealth University 7 January 2013
2 Biological Roles of GAGs Growth Factors cell proliferation, etc. Cytokines PF 4; IL 8; Inflammation Antithrombin; CII anticoagulation eparin / S Dermatan sulfate gb, gc, gd; gp120 viral invasion Annexin V anticoagulation C1 inhibitor complement activation 2
3 Glycosaminoglycans (GAGs) Linear (i.e. unbranched) chains consisting of repeating disaccharide units variously substituted with acetyl and sulfate groups. Free in solution or covalently linked to proteins (proteoglycans). Are phenomenally diverse with regard to saccharide ring conformation and substitution pattern. Unlike other biopolymers, there is no genetic code that determines the precise GAG. 3
4 Monosaccharides Aldehydes and react with alcohols to give hemiacetals R + R' R' R Glucose: an aldose C C 3 1 C 5 6C 2 FISCER Projection C 3 6 C C 3 6C anomeric carbon 6C D-glucopyranose ( -D-glucose) -D-glucopyranose ( -D-glucose) AWRT Projections Anomer: a stereoisomer that differs in configuration only at the anomeric carbon What is an epimer? 4
5 Pyranose Ring Conformations Based on the Cremer Pople ring puckering indices Q, θ and ψ. The low energy chair (C), boat (B) and skew boat (S) conformations are designated by two numbers corresponding to ring atoms that lie above and below a plane defined by the remaining four ring atoms C 4 4 C 1 The atom above the plane is superscripted; the one below is subscripted. 2 S 2 The energetically most favorable conformations for GAGs are 1 C 4, 4 C 1 and 2 S. Fushinobu, S. et al. Carbohydr. Res. 2008, 343,
6 Pyranose Ring Conformations -D-glucose 4 C 1 1 C 4 -L-iduronate 1 C 4 C S Courtois, J. Curr. pin. Microbiol. 2009, 12, Lei, P. S. et al. Bioorg. Med. Chem. 1998, 6,
7 Disaccharides GAGs are heterogeneous, polydisperse, acidic polysaccharides 7
8 Glycosidic Linkage The glycosidic linkage consists of two rotatable bonds designated φ and ψ Consistent and regular values of φ and ψ give rise to helical structure for some GAGs. This can be used as a constraint to limit the number of rotatable bonds during the docking procedure. Courtois, J. Curr. pin. Microbiol. 2009, 12,
9 DS elical Allomorphs Mitra, A. K. et al. J. Mol. Biol. 1983, 169,
10 GAG Binding Sites A typical GAG binding site will have the following characteristics: Surface exposed Shallow ighly positively charged (usually with many positively charged amino acid residues that interact with the negatively charged acidic GAG carboxylate and sulfate groups). Thrombin (PDB ID = 1TB6) 10
11 Now we know something about the protein and ligand so what drives the interaction between the two? 11
12 Thermodynamics The affinity of a ligand for a receptor is determined by the free energy of binding ΔG: ΔG = Δ TΔS = RTlnK eq 12
13 Thermodynamics: Enthalpy (Δ) The change in enthalpy is determined by the formation and breakage of inter and intramolecular chemical bonds. For the most part, we need only be concerned with the weak non covalent interactions. owever, there are exceptions (affinity labels and irreversible suicide inhibitors, for example). Bonds that are formed when the ligand binds to the receptor will stabilize (and thus favor) the ligand receptor complex; bonds that are broken when the ligand binds will disfavor the formation of the complex. 13
14 Thermodynamics: Entropy (ΔS) Entropy represents the amount of uncertainty or degrees of freedom in a closed thermodynamic system. In terms of biological molecular interactions, there are two main factors that account for this: Loss of translational and rotational degrees of freedom: The position of the ligand, as well as freely rotating single bonds within the ligand, are more or less fixed in place by the receptor upon binding, resulting in a decrease in entropy that disfavors the formation of the receptor ligand complex. ydrophobic effect: rganized water molecules surrounding the hydrophobic portions of the ligand and possibly inside the binding site are displaced during the docking event, resulting in an increase in entropy that favors the formation of the receptor ligand complex. 14
15 So how do we translate these ideas into something practically useful? 15
16 Molecular Mechanics The very basic idea is to treat molecules as physical entities that are governed by the conventional (nonquantum) Newtonian Laws of Physics. E TTAL = Σ E stretch + Σ E bend + Σ E torsion + Σ E o-o-p + Σ E vdw + Σ E Coulombic The total internal energy of a molecule is a sum of various energy terms representing various stresses and strains on it. 16
17 Molecular Mechanics: E stretch All bonds E stretch = Σ k i (Δr i ) 2 / 2 i=1 (ooke s Law) 17
18 Molecular Mechanics: E bend All angles E bend = Σ k j (Δθ j ) 2 / 2 j=1 18
19 Molecular Mechanics: E torsion All torsions E torsion = Σ V k [1 + S k cos ( n k - ϕ k )] / 2 k=1 19
20 Molecular Mechanics: E o o p d All out-of-planes E o-o-p = Σ k l (Δd l ) 2 / 2 k=1 20
21 Molecular Mechanics: E vdw All nonbonded a 12 ij a 6 ij E vdw = ΣΣφ ij [ - ] i=1 j>1 21
22 Molecular Mechanics: E Coulombic All nonbonded E Coulombic = C ΣΣ Q i Q j i=1 j>1 ε ij r ij Charles Augustin de Coulomb Where do the charges on atoms come from? 22
23 Molecular Mechanics: Atomic Charge Start with electronegativity 23
24 Molecular Mechanics: Atomic Charge calculate bonding (and atomic polarizability) 24
25 Molecular Mechanics: Atomic Charge partial charge on each atom is estimated based on this polarization: In benzene every C has the same electronegativity; thus, all Cs have same partial charge. The s have same charge with opposite sign. The C partial charge reaches a maximum at the meta position relative to other Cs due to the e - -withdrawing methoxycarbonyl group. Amines are e - -releasing and C charge reaches a maximum (relative to the other Cs) at ortho and para. 25
26 We have the basic equations Now we can bring in a computer to do the calculations. 26
27 Molecular Mechanics: Atomic Charge Together these equations comprise a force field that describes how to calculate the potential energy of a molecule. Bond lengths, bond angles and atomic radii are taken from high resolution X ray structures of small molecules and biomacromolecules. The force constants for ooke s law (stretches and bends) are taken from IR and similar experiments. 27
28 Molecular Mechanics: Minimization Energy Minimization Structure ptimization Simultaneous Process By definition the most optimal structure is the one of lowest energy. In vacuum, at 0 K. Although there are a variety of permutations on the algorithms mostly to increase speed the basic idea is: if the energy is decreasing, then the structure is improving. owever, it is not so simple 28
29 Molecular Mechanics: Minimization Start planar Minimum Start boat Local Minimum Global Minimum chair 29
30 Molecular Mechanics: Local Minima Exhaustive Search Define structures and calculate energies for all conformers. Structure with lowest energy is at Global minimum. The process is generally varying torsion angles for rotatable bonds. At a 15 o increment, number of structures = 24 n, where n is number of these bonds. Molecular Dynamics Simply, add heat and time to molecular mechanics and break out of local minima. Use with larger molecules. 30
31 Molecular Mechanics: Rotatable Bonds GlcN2Ac6S ( 4 C 1 ) IdoA ( 2 S ) 7 to 15 rotatable bonds depending on how you count them. Becomes an issue when docking larger oligosaccharides. Can use the rigid backbone approximation to partially address this. 31
32 Molecular Dynamics 32
33 Docking Involves finding an optimal complementary fit between a ligand molecule and a suitable binding site in a receptor, and typically involves three operations: Generate plausible candidate ligand conformations In general, the conformation(s) that the ligand adopts when bound to the receptor (i.e. the binding mode) is not known. The docking program must therefore generate a number of possible conformations to test. Place the ligand into the binding site The docking program must be able to place the ligand into the binding site such that the distances and angles between complementary functional groups are chemically meaningful and that the atoms are not clashing with one another. Assign a score or fitness value to the docked conformation. Ligand docking is treated computationally as a mathematical optimization problem. Thus, the docked solutions (or poses) that are generated must be evaluated by a fitness or scoring function to determine the quality of each pose. 33
34 GAG Virtual Libraries GAG virtual libraries capture the phenomenal diversity of GAGs by enumerating the sulfation and acetylation patterns of the GAG sequences, as well as the GAG conformations at the monosaccharide level. Example: /S hexasaccharide library: o 13 naturally occurring IdoAp containing disaccharides 2 conformations per IdoAp = 26 IdoAp disaccharides + 10 naturally occurring GlcAp containing disaccharides = 36 virtual /S disaccharides. o 1 hexasaccharide sequence = 3 disaccharide units o = 36 3 = 46,656 virtual /S sequences Esko, J. D.; Selleck, S. B. Annu. Rev. Biochem. 2002, 71,
35 Virtual Screening Virtual screening (VS) provides a means to dock many GAG sequences to a given protein and select the ones that bind with high affinity and selectivity. For large libraries, a dual filter strategy is used : Start Build Virtual Library of /S sequences (Use Average Backbone Model) I st GLD Docking (10,000 iterations; one run per sequence) Affinity Filter (GLD score in top ~1%) N YES II nd GLD Docking (100,000 iterations; three runs per sequence) Specificity Filter (RMSD < 2.5 Å) YES igh Affinity & igh Specificity Sequence(s) N Discard Sidhu, P. S.; Mosier, P. D.; Zhou, Q.; Desai, U. R. Bioorg. Med. Chem. Lett. 2013, 23,
36 Modeling GAG Protein Interactions Molecular modeling provides us with the tools to model ANY GAG protein interaction. Powerful and easy and to do, but filled with caveats: Flexibility of the GAG and/or binding site amino acid residues Ionization state of charged groups Involvement of water 36
37 T AT Binding Site Analysis Identification of highly relevant water molecules Displacement of these water molecules greatly enhances the affinity of the GAG for its binding site. antithrombin Mosier, P. D.; Krishnasamy, C.; Kellogg, G. E.; Desai, U. R. PLoS NE 2012, 7, e
38 Case Study: S AT Interaction The /S pentasaccharide sequence (i.e. DEFG ) binds with with high specificity to AT. Screening of a combinatorial virtual library of 6,859 heparin hexasaccharides using a dual filter strategy, in which antithrombin affinity was the first filter and self consistency of docking was the second, resulted in only 10 sequences. The results explain the biochemical observations with single substitution and ring truncation variants of the pentasaccharide. Raghuraman, A.; Mosier, P. D.; Desai, U. R. J. Med. Chem. 2006, 49,
39 Case Study: S AT Interaction X-ray GLD D E F G 6AE 2S F 3S F K114 R129 6S D K125 ha hp hd Raghuraman, A.; Mosier, P. D.; Desai, U. R. J. Med. Chem. 2006, 49,
40 Case Study: S AT Interaction A) 6A E 2S F 3S F 6S D D E F G D E F G B) Raghuraman, A.; Mosier, P. D.; Desai, U. R. J. Med. Chem. 2006, 49,
41 Case Study: S AT Interaction D E F G C 2 V _ C C 2 S 3 C 2 S 3 _ C W X Y _ NS 3 NS 3 S 3 Z NZ' Me V W X Y Z Z 5+3S S 3 S 3 S 3 S S D S 3 S S F S 3 S S S 3 S 3 5+3S E S 3 S 3 S 3 S S G S 3 S 3 S 3 S - 3 _ C 2 S 3 C C 2 S 3 C 5- S 3 Me NS 3 NS 3 _ C 2 S 3 C C 2 S 3 5-G NS 3 S 3 _ Me NS 3 _ C C 2 S 3 C 2 S 3 Gold Score /K 114 5/K D 5-DE 5-5-G 5 5+3S 5/R G BS (kcal/mol) 5-D C S 3 _ NS 3 _ C 2 S 3 _ S 3 _ S 3 _ Me S 3 _ C 2 S 3 5-DE _ S 3 Me _ S 3 _ C _ S 3 _ S 3 Me Raghuraman, A.; Mosier, P. D.; Desai, U. R. J. Med. Chem. 2006, 49,
42 Conclusions Molecular modeling techniques provide powerful and general methods to address GAG protein binding. These tools can be used to identify potential highaffinity, high specificity GAG sequences for ANY potential GAG binding site. 42
43 Acknowledgments VCU Dr. Umesh Desai Dr. Glen Kellogg All of you! NI P01 L
Carbohydrate- Protein interac;ons are Cri;cal in Life and Death. Other Cells. Hormones. Viruses. Toxins. Cell. Bacteria
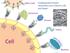 ther Cells Carbohydrate- Protein interac;ons are Cri;cal in Life and Death ormones Viruses Toxins Cell Bacteria ow to Model Protein- ligand interac;ons? Protein Protein Protein DNA/RNA Protein Carbohydrate
ther Cells Carbohydrate- Protein interac;ons are Cri;cal in Life and Death ormones Viruses Toxins Cell Bacteria ow to Model Protein- ligand interac;ons? Protein Protein Protein DNA/RNA Protein Carbohydrate
Other Cells. Hormones. Viruses. Toxins. Cell. Bacteria
 Other Cells Hormones Viruses Toxins Cell Bacteria ΔH < 0 reaction is exothermic, tells us nothing about the spontaneity of the reaction Δ H > 0 reaction is endothermic, tells us nothing about the spontaneity
Other Cells Hormones Viruses Toxins Cell Bacteria ΔH < 0 reaction is exothermic, tells us nothing about the spontaneity of the reaction Δ H > 0 reaction is endothermic, tells us nothing about the spontaneity
Kd = koff/kon = [R][L]/[RL]
![Kd = koff/kon = [R][L]/[RL] Kd = koff/kon = [R][L]/[RL]](/thumbs/96/127564193.jpg) Taller de docking y cribado virtual: Uso de herramientas computacionales en el diseño de fármacos Docking program GLIDE El programa de docking GLIDE Sonsoles Martín-Santamaría Shrödinger is a scientific
Taller de docking y cribado virtual: Uso de herramientas computacionales en el diseño de fármacos Docking program GLIDE El programa de docking GLIDE Sonsoles Martín-Santamaría Shrödinger is a scientific
Loudon Chapter 24 Review: Carbohydrates Jacquie Richardson, CU Boulder Last updated 4/26/2018
 This chapter is about carbohydrates molecules with the general formula of C n( 2O) n, or in other words C n 2nO n. This is a very common formula for sugars and many other natural products. The structure
This chapter is about carbohydrates molecules with the general formula of C n( 2O) n, or in other words C n 2nO n. This is a very common formula for sugars and many other natural products. The structure
User Guide for LeDock
 User Guide for LeDock Hongtao Zhao, PhD Email: htzhao@lephar.com Website: www.lephar.com Copyright 2017 Hongtao Zhao. All rights reserved. Introduction LeDock is flexible small-molecule docking software,
User Guide for LeDock Hongtao Zhao, PhD Email: htzhao@lephar.com Website: www.lephar.com Copyright 2017 Hongtao Zhao. All rights reserved. Introduction LeDock is flexible small-molecule docking software,
Using Bayesian Statistics to Predict Water Affinity and Behavior in Protein Binding Sites. J. Andrew Surface
 Using Bayesian Statistics to Predict Water Affinity and Behavior in Protein Binding Sites Introduction J. Andrew Surface Hampden-Sydney College / Virginia Commonwealth University In the past several decades
Using Bayesian Statistics to Predict Water Affinity and Behavior in Protein Binding Sites Introduction J. Andrew Surface Hampden-Sydney College / Virginia Commonwealth University In the past several decades
Docking. GBCB 5874: Problem Solving in GBCB
 Docking Benzamidine Docking to Trypsin Relationship to Drug Design Ligand-based design QSAR Pharmacophore modeling Can be done without 3-D structure of protein Receptor/Structure-based design Molecular
Docking Benzamidine Docking to Trypsin Relationship to Drug Design Ligand-based design QSAR Pharmacophore modeling Can be done without 3-D structure of protein Receptor/Structure-based design Molecular
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Loudon Chapter 24 Review: Carbohydrates CHEM 3331, Jacquie Richardson, Fall Page 1
 CEM 3331, Jacquie Richardson, Fall 2010 - Page 1 This chapter is about carbohydrates molecules with the general formula of C n ( 2 ) n, or in other words C n 2n n. This is a very common formula for sugars
CEM 3331, Jacquie Richardson, Fall 2010 - Page 1 This chapter is about carbohydrates molecules with the general formula of C n ( 2 ) n, or in other words C n 2n n. This is a very common formula for sugars
Protein-Ligand Interactions* and Energy Evaluation Methods
 Protein-Ligand Interactions* and Energy Evaluation Methods *with a revealing look at roles of water Glen E. Kellogg Department of Medicinal Chemistry Institute for Structural Biology, Drug Discovery &
Protein-Ligand Interactions* and Energy Evaluation Methods *with a revealing look at roles of water Glen E. Kellogg Department of Medicinal Chemistry Institute for Structural Biology, Drug Discovery &
ORGANIC - BROWN 8E CH CARBOHYDRATES.
 !! www.clutchprep.com CONCEPT: INTRODUCTION TO CARBOHYDRATE MONOSACCHARIDES Sugars or saccharides are also referred to as carbo-hydrates, implying that carbon has been combined with Monosaccharides are
!! www.clutchprep.com CONCEPT: INTRODUCTION TO CARBOHYDRATE MONOSACCHARIDES Sugars or saccharides are also referred to as carbo-hydrates, implying that carbon has been combined with Monosaccharides are
Structural Bioinformatics (C3210) Molecular Docking
 Structural Bioinformatics (C3210) Molecular Docking Molecular Recognition, Molecular Docking Molecular recognition is the ability of biomolecules to recognize other biomolecules and selectively interact
Structural Bioinformatics (C3210) Molecular Docking Molecular Recognition, Molecular Docking Molecular recognition is the ability of biomolecules to recognize other biomolecules and selectively interact
BCMP 201 Protein biochemistry
 BCMP 201 Protein biochemistry BCMP 201 Protein biochemistry with emphasis on the interrelated roles of protein structure, catalytic activity, and macromolecular interactions in biological processes. The
BCMP 201 Protein biochemistry BCMP 201 Protein biochemistry with emphasis on the interrelated roles of protein structure, catalytic activity, and macromolecular interactions in biological processes. The
25.6 Cyclic Forms of Carbohydrates: Furanose Forms
 .6 Cyclic Forms of Carbohydrates: Furanose Forms Recall from Section 7.8 R R' C + R" Product is a hemiacetal. R" C R R' R C Cyclic emiacetals R C Aldehydes and ketones that contain an group elsewhere in
.6 Cyclic Forms of Carbohydrates: Furanose Forms Recall from Section 7.8 R R' C + R" Product is a hemiacetal. R" C R R' R C Cyclic emiacetals R C Aldehydes and ketones that contain an group elsewhere in
25.3 Cyclization of Monosaccharides
 090 CAPTER CARBYDRATES PRBLEM. Determine the identity of each of these carbohydrates: C a) b) c) C C C C C (int: This must be rotated first.). Cyclization of Monosaccharides Up to this point, the structure
090 CAPTER CARBYDRATES PRBLEM. Determine the identity of each of these carbohydrates: C a) b) c) C C C C C (int: This must be rotated first.). Cyclization of Monosaccharides Up to this point, the structure
Section Week 3. Junaid Malek, M.D.
 Section Week 3 Junaid Malek, M.D. Biological Polymers DA 4 monomers (building blocks), limited structure (double-helix) RA 4 monomers, greater flexibility, multiple structures Proteins 20 Amino Acids,
Section Week 3 Junaid Malek, M.D. Biological Polymers DA 4 monomers (building blocks), limited structure (double-helix) RA 4 monomers, greater flexibility, multiple structures Proteins 20 Amino Acids,
Molecular Mechanics, Dynamics & Docking
 Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
Solutions and Non-Covalent Binding Forces
 Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
ORGANIC - BRUICE 8E CH THE ORGANIC CHEMISTRY OF CARBOHYDRATES
 !! www.clutchprep.com CONCEPT: INTRODUCTION TO CARBOHYDRATE MONOSACCHARIDES Sugars or saccharides are also referred to as carbo-hydrates, implying that carbon has been combined with Monosaccharides are
!! www.clutchprep.com CONCEPT: INTRODUCTION TO CARBOHYDRATE MONOSACCHARIDES Sugars or saccharides are also referred to as carbo-hydrates, implying that carbon has been combined with Monosaccharides are
NMR solution conformation of heparin-derived hexasaccharide
 Biochem. J. (1997) 328, 51 61 (Printed in Great Britain) 51 NMR solution conformation of heparin-derived hexasaccharide Dmitri MIKHAILOV*, Robert J. LINHARDT and Kevin H. MAYO* 1 *Departments of Biochemistry,
Biochem. J. (1997) 328, 51 61 (Printed in Great Britain) 51 NMR solution conformation of heparin-derived hexasaccharide Dmitri MIKHAILOV*, Robert J. LINHARDT and Kevin H. MAYO* 1 *Departments of Biochemistry,
Biomolecules. Energetics in biology. Biomolecules inside the cell
 Biomolecules Energetics in biology Biomolecules inside the cell Energetics in biology The production of energy, its storage, and its use are central to the economy of the cell. Energy may be defined as
Biomolecules Energetics in biology Biomolecules inside the cell Energetics in biology The production of energy, its storage, and its use are central to the economy of the cell. Energy may be defined as
CHAPTER 3. Vibrational Characteristics of PTP-1B Inhibitors
 CHAPTER 3 Vibrational Characteristics of PTP-1B Inhibitors 3.1 Preface Theoretical Frequency calculations were performed on the molecules chosen in this study. Initially all the geometries were optimized
CHAPTER 3 Vibrational Characteristics of PTP-1B Inhibitors 3.1 Preface Theoretical Frequency calculations were performed on the molecules chosen in this study. Initially all the geometries were optimized
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability
 Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Chapter 1. Topic: Overview of basic principles
 Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions Van der Waals Interactions
Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions Van der Waals Interactions
Cholera Toxin Invasion
 Protein-carbohydrate interactions: Isothermal Titration Calorimetry Dr Bruce Turnbull School of Chemistry and Astbury Centre for Structural Molecular Biology University of Leeds Cholera Toxin Invasion
Protein-carbohydrate interactions: Isothermal Titration Calorimetry Dr Bruce Turnbull School of Chemistry and Astbury Centre for Structural Molecular Biology University of Leeds Cholera Toxin Invasion
ORGANIC - CLUTCH CH CARBOHYDRATES.
 !! www.clutchprep.com CONCEPT: INTRODUCTION TO CARBOHYDRATE MONOSACCHARIDES Sugars or saccharides are also referred to as carbo-hydrates, implying that carbon has been combined with Monosaccharides are
!! www.clutchprep.com CONCEPT: INTRODUCTION TO CARBOHYDRATE MONOSACCHARIDES Sugars or saccharides are also referred to as carbo-hydrates, implying that carbon has been combined with Monosaccharides are
= (-22) = +2kJ /mol
 Lecture 8: Thermodynamics & Protein Stability Assigned reading in Campbell: Chapter 4.4-4.6 Key Terms: DG = -RT lnk eq = DH - TDS Transition Curve, Melting Curve, Tm DH calculation DS calculation van der
Lecture 8: Thermodynamics & Protein Stability Assigned reading in Campbell: Chapter 4.4-4.6 Key Terms: DG = -RT lnk eq = DH - TDS Transition Curve, Melting Curve, Tm DH calculation DS calculation van der
11/14/2016. NaBH 4. Superimposable on mirror image. (achiral) D-ribose. NaBH 4 NaBH 4. Not superimposable on mirror image.
 There are multiple ways to relate the structures of ketoses to aldoses. Reduction of a ketose with NaB 4 forms a new stereocenter and thus forms a mixture of two alditols. An aldopentose forms a chiral
There are multiple ways to relate the structures of ketoses to aldoses. Reduction of a ketose with NaB 4 forms a new stereocenter and thus forms a mixture of two alditols. An aldopentose forms a chiral
Chemical properties that affect binding of enzyme-inhibiting drugs to enzymes
 Chemical properties that affect binding of enzyme-inhibiting drugs to enzymes Introduction The production of new drugs requires time for development and testing, and can result in large prohibitive costs
Chemical properties that affect binding of enzyme-inhibiting drugs to enzymes Introduction The production of new drugs requires time for development and testing, and can result in large prohibitive costs
Homework Problem Set 4 Solutions
 Chemistry 380.37 Dr. Jean M. Standard omework Problem Set 4 Solutions 1. A conformation search is carried out on a system and four low energy stable conformers are obtained. Using the MMFF force field,
Chemistry 380.37 Dr. Jean M. Standard omework Problem Set 4 Solutions 1. A conformation search is carried out on a system and four low energy stable conformers are obtained. Using the MMFF force field,
Protein-Ligand Docking Methods
 Review Goal: Given a protein structure, predict its ligand bindings Protein-Ligand Docking Methods Applications: Function prediction Drug discovery etc. Thomas Funkhouser Princeton University S597A, Fall
Review Goal: Given a protein structure, predict its ligand bindings Protein-Ligand Docking Methods Applications: Function prediction Drug discovery etc. Thomas Funkhouser Princeton University S597A, Fall
THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION
 THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION AND CALIBRATION Calculation of turn and beta intrinsic propensities. A statistical analysis of a protein structure
THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION AND CALIBRATION Calculation of turn and beta intrinsic propensities. A statistical analysis of a protein structure
Question 1. Identify the sugars below by filling in the table below (except shaded areas). Use the page or a separate sheet
 Question 1. Identify the sugars below by filling in the table below (except shaded areas). Use the page or a separate sheet Sugar name aworth projection(s) of the corresponding pyranose form (Six membered
Question 1. Identify the sugars below by filling in the table below (except shaded areas). Use the page or a separate sheet Sugar name aworth projection(s) of the corresponding pyranose form (Six membered
Importance of Carbohydrates
 Chapter 25 Importance of Carbohydrates Distributed widely in nature Key intermediates of metabolism (sugars) Structural components of plants (cellulose) Central to materials of industrial products: paper,
Chapter 25 Importance of Carbohydrates Distributed widely in nature Key intermediates of metabolism (sugars) Structural components of plants (cellulose) Central to materials of industrial products: paper,
Chap. 3 Conformational Analysis and Molecular Mechanics
 hap. 3 onformational Analysis and Molecular Mechanics onformation:ne of several different spatial arrangements that a molecule can achieve by rotation about single bonds between atoms. Notations: -sc -ac
hap. 3 onformational Analysis and Molecular Mechanics onformation:ne of several different spatial arrangements that a molecule can achieve by rotation about single bonds between atoms. Notations: -sc -ac
Structural biology and drug design: An overview
 Structural biology and drug design: An overview livier Taboureau Assitant professor Chemoinformatics group-cbs-dtu otab@cbs.dtu.dk Drug discovery Drug and drug design A drug is a key molecule involved
Structural biology and drug design: An overview livier Taboureau Assitant professor Chemoinformatics group-cbs-dtu otab@cbs.dtu.dk Drug discovery Drug and drug design A drug is a key molecule involved
The PhilOEsophy. There are only two fundamental molecular descriptors
 The PhilOEsophy There are only two fundamental molecular descriptors Where can we use shape? Virtual screening More effective than 2D Lead-hopping Shape analogues are not graph analogues Molecular alignment
The PhilOEsophy There are only two fundamental molecular descriptors Where can we use shape? Virtual screening More effective than 2D Lead-hopping Shape analogues are not graph analogues Molecular alignment
Chemical properties that affect binding of enzyme-inhibiting drugs to enzymes
 Introduction Chemical properties that affect binding of enzyme-inhibiting drugs to enzymes The production of new drugs requires time for development and testing, and can result in large prohibitive costs
Introduction Chemical properties that affect binding of enzyme-inhibiting drugs to enzymes The production of new drugs requires time for development and testing, and can result in large prohibitive costs
A. Reaction Mechanisms and Catalysis (1) proximity effect (2) acid-base catalysts (3) electrostatic (4) functional groups (5) structural flexibility
 (P&S Ch 5; Fer Ch 2, 9; Palm Ch 10,11; Zub Ch 9) A. Reaction Mechanisms and Catalysis (1) proximity effect (2) acid-base catalysts (3) electrostatic (4) functional groups (5) structural flexibility B.
(P&S Ch 5; Fer Ch 2, 9; Palm Ch 10,11; Zub Ch 9) A. Reaction Mechanisms and Catalysis (1) proximity effect (2) acid-base catalysts (3) electrostatic (4) functional groups (5) structural flexibility B.
T6.2 Molecular Mechanics
 T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
Protein-Ligand Docking Methods
 Review Goal: Given a protein structure, predict its ligand bindings Protein-Ligand Docking Methods Applications: Function prediction Drug discovery etc. Thomas Funkhouser Princeton University S597A, Fall
Review Goal: Given a protein structure, predict its ligand bindings Protein-Ligand Docking Methods Applications: Function prediction Drug discovery etc. Thomas Funkhouser Princeton University S597A, Fall
Chem 204. Mid-Term Exam I. July 21, There are 3 sections to this exam: Answer ALL questions
 Chem 204 Mid-Term Exam I July 21, 2009 Name: Answer Key Student ID: There are 3 sections to this exam: Answer ALL questions Section I: Multiple-Choice 20 questions, 2 pts each Section II: Fill-in-the-Blank
Chem 204 Mid-Term Exam I July 21, 2009 Name: Answer Key Student ID: There are 3 sections to this exam: Answer ALL questions Section I: Multiple-Choice 20 questions, 2 pts each Section II: Fill-in-the-Blank
BIBC 100. Structural Biochemistry
 BIBC 100 Structural Biochemistry http://classes.biology.ucsd.edu/bibc100.wi14 Papers- Dialogue with Scientists Questions: Why? How? What? So What? Dialogue Structure to explain function Knowledge Food
BIBC 100 Structural Biochemistry http://classes.biology.ucsd.edu/bibc100.wi14 Papers- Dialogue with Scientists Questions: Why? How? What? So What? Dialogue Structure to explain function Knowledge Food
Fondamenti di Chimica Farmaceutica. Computer Chemistry in Drug Research: Introduction
 Fondamenti di Chimica Farmaceutica Computer Chemistry in Drug Research: Introduction Introduction Introduction Introduction Computer Chemistry in Drug Design Drug Discovery: Target identification Lead
Fondamenti di Chimica Farmaceutica Computer Chemistry in Drug Research: Introduction Introduction Introduction Introduction Computer Chemistry in Drug Design Drug Discovery: Target identification Lead
High Throughput In-Silico Screening Against Flexible Protein Receptors
 John von Neumann Institute for Computing High Throughput In-Silico Screening Against Flexible Protein Receptors H. Sánchez, B. Fischer, H. Merlitz, W. Wenzel published in From Computational Biophysics
John von Neumann Institute for Computing High Throughput In-Silico Screening Against Flexible Protein Receptors H. Sánchez, B. Fischer, H. Merlitz, W. Wenzel published in From Computational Biophysics
`1AP Biology Study Guide Chapter 2 v Atomic structure is the basis of life s chemistry Ø Living and non- living things are composed of atoms Ø
 `1AP Biology Study Guide Chapter 2 v Atomic structure is the basis of life s chemistry Ø Living and non- living things are composed of atoms Ø Element pure substance only one kind of atom Ø Living things
`1AP Biology Study Guide Chapter 2 v Atomic structure is the basis of life s chemistry Ø Living and non- living things are composed of atoms Ø Element pure substance only one kind of atom Ø Living things
ORGANIC - EGE 5E EXTRA: CARBOHYDRATES.
 !! www.clutchprep.com CONCEPT: INTRODUCTION TO CARBOHYDRATE MONOSACCHARIDES Sugars or saccharides are also referred to as carbo-hydrates, implying that carbon has been combined with Monosaccharides are
!! www.clutchprep.com CONCEPT: INTRODUCTION TO CARBOHYDRATE MONOSACCHARIDES Sugars or saccharides are also referred to as carbo-hydrates, implying that carbon has been combined with Monosaccharides are
Protein Structure. W. M. Grogan, Ph.D. OBJECTIVES
 Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
Carbohydrates - Introduction
 arbohydrates - Introduction arbohydrates - General Description A. Polyhydroxy Aldehydes or Ketones ARBN AIN B. Serve a variety of functions ARBN AIN ARBN AIN 1. Energy storage (Glucose, Glycogen, Starch)
arbohydrates - Introduction arbohydrates - General Description A. Polyhydroxy Aldehydes or Ketones ARBN AIN B. Serve a variety of functions ARBN AIN ARBN AIN 1. Energy storage (Glucose, Glycogen, Starch)
DISCRETE TUTORIAL. Agustí Emperador. Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING:
 DISCRETE TUTORIAL Agustí Emperador Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING: STRUCTURAL REFINEMENT OF DOCKING CONFORMATIONS Emperador
DISCRETE TUTORIAL Agustí Emperador Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING: STRUCTURAL REFINEMENT OF DOCKING CONFORMATIONS Emperador
Dihedral Angles. Homayoun Valafar. Department of Computer Science and Engineering, USC 02/03/10 CSCE 769
 Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
Chemistry in Biology. Section 1. Atoms, Elements, and Compounds
 Section 1 Atoms, Elements, and Compounds Atoms! Chemistry is the study of matter.! Atoms are the building blocks of matter.! Neutrons and protons are located at the center of the atom.! Protons are positively
Section 1 Atoms, Elements, and Compounds Atoms! Chemistry is the study of matter.! Atoms are the building blocks of matter.! Neutrons and protons are located at the center of the atom.! Protons are positively
Free energy, electrostatics, and the hydrophobic effect
 Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
Assignment 1: Molecular Mechanics (PART 1 25 points)
 Chemistry 380.37 Fall 2015 Dr. Jean M. Standard August 19, 2015 Assignment 1: Molecular Mechanics (PART 1 25 points) In this assignment, you will perform some molecular mechanics calculations using the
Chemistry 380.37 Fall 2015 Dr. Jean M. Standard August 19, 2015 Assignment 1: Molecular Mechanics (PART 1 25 points) In this assignment, you will perform some molecular mechanics calculations using the
Examples of Protein Modeling. Protein Modeling. Primary Structure. Protein Structure Description. Protein Sequence Sources. Importing Sequences to MOE
 Examples of Protein Modeling Protein Modeling Visualization Examination of an experimental structure to gain insight about a research question Dynamics To examine the dynamics of protein structures To
Examples of Protein Modeling Protein Modeling Visualization Examination of an experimental structure to gain insight about a research question Dynamics To examine the dynamics of protein structures To
The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii
 The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii Fundamental Points About Which to Be Thinking It s clear the PES is useful, so how can I construct it for an arbitrary
The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii Fundamental Points About Which to Be Thinking It s clear the PES is useful, so how can I construct it for an arbitrary
Softwares for Molecular Docking. Lokesh P. Tripathi NCBS 17 December 2007
 Softwares for Molecular Docking Lokesh P. Tripathi NCBS 17 December 2007 Molecular Docking Attempt to predict structures of an intermolecular complex between two or more molecules Receptor-ligand (or drug)
Softwares for Molecular Docking Lokesh P. Tripathi NCBS 17 December 2007 Molecular Docking Attempt to predict structures of an intermolecular complex between two or more molecules Receptor-ligand (or drug)
Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation
 Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation Jakob P. Ulmschneider and William L. Jorgensen J.A.C.S. 2004, 126, 1849-1857 Presented by Laura L. Thomas and
Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation Jakob P. Ulmschneider and William L. Jorgensen J.A.C.S. 2004, 126, 1849-1857 Presented by Laura L. Thomas and
Protein Dynamics. The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron.
 Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Life Sciences 1a Lecture Slides Set 10 Fall Prof. David R. Liu. Lecture Readings. Required: Lecture Notes McMurray p , O NH
 Life ciences 1a Lecture lides et 10 Fall 2006-2007 Prof. David R. Liu Lectures 17-18: The molecular basis of drug-protein binding: IV protease inhibitors 1. Drug development and its impact on IV-infected
Life ciences 1a Lecture lides et 10 Fall 2006-2007 Prof. David R. Liu Lectures 17-18: The molecular basis of drug-protein binding: IV protease inhibitors 1. Drug development and its impact on IV-infected
Electronic Supplementary Information Effective lead optimization targeted for displacing bridging water molecule
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Effective lead optimization targeted for displacing
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Effective lead optimization targeted for displacing
What is Protein-Ligand Docking?
 MOLECULAR DOCKING Definition: What is Protein-Ligand Docking? Computationally predict the structures of protein-ligand complexes from their conformations and orientations. The orientation that maximizes
MOLECULAR DOCKING Definition: What is Protein-Ligand Docking? Computationally predict the structures of protein-ligand complexes from their conformations and orientations. The orientation that maximizes
Microcalorimetry for the Life Sciences
 Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Why Proteins Fold. How Proteins Fold? e - ΔG/kT. Protein Folding, Nonbonding Forces, and Free Energy
 Why Proteins Fold Proteins are the action superheroes of the body. As enzymes, they make reactions go a million times faster. As versatile transport vehicles, they carry oxygen and antibodies to fight
Why Proteins Fold Proteins are the action superheroes of the body. As enzymes, they make reactions go a million times faster. As versatile transport vehicles, they carry oxygen and antibodies to fight
Bio-elements. Living organisms requires only 27 of the 90 common chemical elements found in the crust of the earth, to be as its essential components.
 Bio-elements Living organisms requires only 27 of the 90 common chemical elements found in the crust of the earth, to be as its essential components. Most of the chemical components of living organisms
Bio-elements Living organisms requires only 27 of the 90 common chemical elements found in the crust of the earth, to be as its essential components. Most of the chemical components of living organisms
The biomolecules of terrestrial life
 Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Chapter 24 Carbohydrates
 hapter arbohydrates Review of oncepts Fill in the blanks below. To verify that your answers are correct, look in your textbook at the end of hapter. Each of the sentences below appears verbatim in the
hapter arbohydrates Review of oncepts Fill in the blanks below. To verify that your answers are correct, look in your textbook at the end of hapter. Each of the sentences below appears verbatim in the
Chapter 25 Organic and Biological Chemistry
 Chapter 25 Organic and Biological Chemistry Organic Chemistry The chemistry of carbon compounds. Carbon has the ability to form long chains. Without this property, large biomolecules such as proteins,
Chapter 25 Organic and Biological Chemistry Organic Chemistry The chemistry of carbon compounds. Carbon has the ability to form long chains. Without this property, large biomolecules such as proteins,
Carbon and Molecular Diversity - 1
 Carbon and Molecular Diversity - 1 Although water is the most abundant compound of living organisms, and the "medium" for the existence of life, most of the molecules from which living organisms are composed
Carbon and Molecular Diversity - 1 Although water is the most abundant compound of living organisms, and the "medium" for the existence of life, most of the molecules from which living organisms are composed
Bioengineering & Bioinformatics Summer Institute, Dept. Computational Biology, University of Pittsburgh, PGH, PA
 Pharmacophore Model Development for the Identification of Novel Acetylcholinesterase Inhibitors Edwin Kamau Dept Chem & Biochem Kennesa State Uni ersit Kennesa GA 30144 Dept. Chem. & Biochem. Kennesaw
Pharmacophore Model Development for the Identification of Novel Acetylcholinesterase Inhibitors Edwin Kamau Dept Chem & Biochem Kennesa State Uni ersit Kennesa GA 30144 Dept. Chem. & Biochem. Kennesaw
Molecular Modeling -- Lecture 15 Surfaces and electrostatics
 Molecular Modeling -- Lecture 15 Surfaces and electrostatics Molecular surfaces The Hydrophobic Effect Electrostatics Poisson-Boltzmann Equation Electrostatic maps Electrostatic surfaces in MOE 15.1 The
Molecular Modeling -- Lecture 15 Surfaces and electrostatics Molecular surfaces The Hydrophobic Effect Electrostatics Poisson-Boltzmann Equation Electrostatic maps Electrostatic surfaces in MOE 15.1 The
The protein folding problem consists of two parts:
 Energetics and kinetics of protein folding The protein folding problem consists of two parts: 1)Creating a stable, well-defined structure that is significantly more stable than all other possible structures.
Energetics and kinetics of protein folding The protein folding problem consists of two parts: 1)Creating a stable, well-defined structure that is significantly more stable than all other possible structures.
Model Worksheet Teacher Key
 Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Protein-Ligand Docking
 Protein-Ligand Docking Matthias Rarey GMD - German National Research Center for Information Technology Institute for Algorithms and Scientific Computing (SCAI) 53754Sankt Augustin, Germany rarey@gmd.de
Protein-Ligand Docking Matthias Rarey GMD - German National Research Center for Information Technology Institute for Algorithms and Scientific Computing (SCAI) 53754Sankt Augustin, Germany rarey@gmd.de
Virtual screening for drug discovery. Markus Lill Purdue University
 Virtual screening for drug discovery Markus Lill Purdue University mlill@purdue.edu Lecture material http://people.pharmacy.purdue.edu/~mlill/teaching/eidelberg/ I.1 Drug discovery Cl N Disease I.1 Drug
Virtual screening for drug discovery Markus Lill Purdue University mlill@purdue.edu Lecture material http://people.pharmacy.purdue.edu/~mlill/teaching/eidelberg/ I.1 Drug discovery Cl N Disease I.1 Drug
Atomic weight = Number of protons + neutrons
 1 BIOLOGY Elements and Compounds Element is a substance that cannot be broken down to other substances by chemical reactions. Essential elements are chemical elements required for an organism to survive,
1 BIOLOGY Elements and Compounds Element is a substance that cannot be broken down to other substances by chemical reactions. Essential elements are chemical elements required for an organism to survive,
Dr. Sander B. Nabuurs. Computational Drug Discovery group Center for Molecular and Biomolecular Informatics Radboud University Medical Centre
 Dr. Sander B. Nabuurs Computational Drug Discovery group Center for Molecular and Biomolecular Informatics Radboud University Medical Centre The road to new drugs. How to find new hits? High Throughput
Dr. Sander B. Nabuurs Computational Drug Discovery group Center for Molecular and Biomolecular Informatics Radboud University Medical Centre The road to new drugs. How to find new hits? High Throughput
Chem 215 F11 Notes Dr. Masato Koreeda - Page 1 of 18. Date: November 9, 2011
 Chem F Notes Dr. Masato Koreeda - Page of 8. Date: November 9, 0 Chapters.8; -,,, and 7: Carbohydrates Part I Carbohydrate nomenclature: http://www.chem.qmul.ac.uk/iupac/carb/ Carbohydrates: Polyhydroxylated
Chem F Notes Dr. Masato Koreeda - Page of 8. Date: November 9, 0 Chapters.8; -,,, and 7: Carbohydrates Part I Carbohydrate nomenclature: http://www.chem.qmul.ac.uk/iupac/carb/ Carbohydrates: Polyhydroxylated
LECTURE 5: TYPES OF CARBOHYRATE LECTURE OUTCOMES
 LECTURE 5: TYPES OF CARBOHYRATE LECTURE OUTCOMES After completing this lecture and mastering the lecture materials, students are expected to be able 1. 2. 3. 4. 5. to explain the origin of plants carbohydrates
LECTURE 5: TYPES OF CARBOHYRATE LECTURE OUTCOMES After completing this lecture and mastering the lecture materials, students are expected to be able 1. 2. 3. 4. 5. to explain the origin of plants carbohydrates
Protein-Ligand Docking Evaluations
 Introduction Protein-Ligand Docking Evaluations Protein-ligand docking: Given a protein and a ligand, determine the pose(s) and conformation(s) minimizing the total energy of the protein-ligand complex
Introduction Protein-Ligand Docking Evaluations Protein-ligand docking: Given a protein and a ligand, determine the pose(s) and conformation(s) minimizing the total energy of the protein-ligand complex
3. Solutions W = N!/(N A!N B!) (3.1) Using Stirling s approximation ln(n!) = NlnN N: ΔS mix = k (N A lnn + N B lnn N A lnn A N B lnn B ) (3.
 3. Solutions Many biological processes occur between molecules in aqueous solution. In addition, many protein and nucleic acid molecules adopt three-dimensional structure ( fold ) in aqueous solution.
3. Solutions Many biological processes occur between molecules in aqueous solution. In addition, many protein and nucleic acid molecules adopt three-dimensional structure ( fold ) in aqueous solution.
MCB100A/Chem130 MidTerm Exam 2 April 4, 2013
 MCBA/Chem Miderm Exam 2 April 4, 2 Name Student ID rue/false (2 points each).. he Boltzmann constant, k b sets the energy scale for observing energy microstates 2. Atoms with favorable electronic configurations
MCBA/Chem Miderm Exam 2 April 4, 2 Name Student ID rue/false (2 points each).. he Boltzmann constant, k b sets the energy scale for observing energy microstates 2. Atoms with favorable electronic configurations
Exam 1 (Monday, July 6, 2015)
 Chem 231 Summer 2015 Assigned Homework Problems Last updated: Friday, July 24, 2015 Problems Assigned from Essential Organic Chemistry, 2 nd Edition, Paula Yurkanis Bruice, Prentice Hall, New York, NY,
Chem 231 Summer 2015 Assigned Homework Problems Last updated: Friday, July 24, 2015 Problems Assigned from Essential Organic Chemistry, 2 nd Edition, Paula Yurkanis Bruice, Prentice Hall, New York, NY,
Ligand-receptor interactions
 University of Silesia, Katowice, Poland 11 22 March 2013 Ligand-receptor interactions Dr. Pavel Polishchuk A.V. Bogatsky Physico-Chemical Institute of National Academy of Sciences of Ukraine Odessa, Ukraine
University of Silesia, Katowice, Poland 11 22 March 2013 Ligand-receptor interactions Dr. Pavel Polishchuk A.V. Bogatsky Physico-Chemical Institute of National Academy of Sciences of Ukraine Odessa, Ukraine
Supporting Information
 Supporting Information Superagonist, Full Agonist, Partial Agonist and Antagonist Actions of Arylguanidines at 5-Hydroxytryptamine-3 (5-HT 3 ) Subunit A Receptors Katie Alix, Shailesh Khatri, Philip D.
Supporting Information Superagonist, Full Agonist, Partial Agonist and Antagonist Actions of Arylguanidines at 5-Hydroxytryptamine-3 (5-HT 3 ) Subunit A Receptors Katie Alix, Shailesh Khatri, Philip D.
The Molecules of Life Chapter 2
 The Molecules of Life Chapter 2 Core concepts 1.The atom is the fundamental unit of matter. 2.Atoms can combine to form molecules linked by chemical bonds. 3.Water is essential for life. 4.Carbon is the
The Molecules of Life Chapter 2 Core concepts 1.The atom is the fundamental unit of matter. 2.Atoms can combine to form molecules linked by chemical bonds. 3.Water is essential for life. 4.Carbon is the
MCB100A/Chem130 MidTerm Exam 2 April 4, 2013
 MCB1A/Chem13 MidTerm Exam 2 April 4, 213 Name Student ID True/False (2 points each). 1. The Boltzmann constant, k b T sets the energy scale for observing energy microstates 2. Atoms with favorable electronic
MCB1A/Chem13 MidTerm Exam 2 April 4, 213 Name Student ID True/False (2 points each). 1. The Boltzmann constant, k b T sets the energy scale for observing energy microstates 2. Atoms with favorable electronic
Chapter 25. Organic and Biological Chemistry. Organic and
 Chapter 25 Calculate grade: (Add exam1 - exam 4 scores)x1.5 Add 7 best quizzes (each quiz is worth 29) Gives you the # points you have so far. Final (worth 200 points) Grades: 800-4 750-3.5 700-3 650-2.5
Chapter 25 Calculate grade: (Add exam1 - exam 4 scores)x1.5 Add 7 best quizzes (each quiz is worth 29) Gives you the # points you have so far. Final (worth 200 points) Grades: 800-4 750-3.5 700-3 650-2.5
Biological Thermodynamics
 Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
F. Piazza Center for Molecular Biophysics and University of Orléans, France. Selected topic in Physical Biology. Lecture 1
 Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Central Dogma. modifications genome transcriptome proteome
 entral Dogma DA ma protein post-translational modifications genome transcriptome proteome 83 ierarchy of Protein Structure 20 Amino Acids There are 20 n possible sequences for a protein of n residues!
entral Dogma DA ma protein post-translational modifications genome transcriptome proteome 83 ierarchy of Protein Structure 20 Amino Acids There are 20 n possible sequences for a protein of n residues!
CHEMISTRY 332 SUMMER 08 EXAM I June 26-28, 2008
 First Three Letters of Last Name NAME Network ID CHEMISTRY 332 SUMMER 08 EXAM I June 26-28, 2008 The following materials are permissible during the exam: molecular model kits, course notes (printed, electronic,
First Three Letters of Last Name NAME Network ID CHEMISTRY 332 SUMMER 08 EXAM I June 26-28, 2008 The following materials are permissible during the exam: molecular model kits, course notes (printed, electronic,
Energy functions and their relationship to molecular conformation. CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror
 Energy functions and their relationship to molecular conformation CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror Yesterday s Nobel Prize: single-particle cryoelectron microscopy 2 Outline Energy
Energy functions and their relationship to molecular conformation CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror Yesterday s Nobel Prize: single-particle cryoelectron microscopy 2 Outline Energy
Water. 2.1 Weak Interactions in Aqueous Sy stems Ionization of Water, Weak Acids, and Weak Bases 58
 Home http://www.macmillanhighered.com/launchpad/lehninger6e... 1 of 1 1/6/2016 3:07 PM 2 Printed Page 47 Water 2.1 Weak Interactions in Aqueous Sy stems 47 2.2 Ionization of Water, Weak Acids, and Weak
Home http://www.macmillanhighered.com/launchpad/lehninger6e... 1 of 1 1/6/2016 3:07 PM 2 Printed Page 47 Water 2.1 Weak Interactions in Aqueous Sy stems 47 2.2 Ionization of Water, Weak Acids, and Weak
O H HO H. !-D-galactopyranose
 ame Key W06-Exam o. Page I. ( points) A disaccharide is cleaved by a β-glycosidase, an enzyme that specifically hydrolyzes a β- glycosidic linkage. When the disaccharide is treated with excess dimethyl
ame Key W06-Exam o. Page I. ( points) A disaccharide is cleaved by a β-glycosidase, an enzyme that specifically hydrolyzes a β- glycosidic linkage. When the disaccharide is treated with excess dimethyl
Protein Structure Prediction and Protein-Ligand Docking
 Protein Structure Prediction and Protein-Ligand Docking Björn Wallner bjornw@ifm.liu.se Jan. 24, 2014 Todays topics Protein Folding Intro Protein structure prediction How can we predict the structure of
Protein Structure Prediction and Protein-Ligand Docking Björn Wallner bjornw@ifm.liu.se Jan. 24, 2014 Todays topics Protein Folding Intro Protein structure prediction How can we predict the structure of
What are the building blocks of life?
 Why? What are the building blocks of life? From the smallest single-celled organism to the tallest tree, all life depends on the properties and reactions of four classes of organic (carbon-based) compounds
Why? What are the building blocks of life? From the smallest single-celled organism to the tallest tree, all life depends on the properties and reactions of four classes of organic (carbon-based) compounds
Structural Bioinformatics (C3210) Molecular Mechanics
 Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
