Late gastrula Early neurula Late neurula
|
|
|
- Rhoda Golden
- 6 years ago
- Views:
Transcription
1 A B Late gastrula Early neurula Late neurula Figure S1 A. The neural crest gene network in vertebrates. Black arrows indicate empirically verified regulatory interactions. In vertebrates, signals from ventral ectoderm and underlying mesendoderm pattern the dorsal ectoderm, inducing expression of neural border specifiers and neural plate markers. These inductive signals then work with neural border specifiers to activate expression of neural crest specifiers. The neural crest specifiers cross-regulate and activate various effector genes, each of which mediates a different aspect of the neural crest phenotype. These include cassettes controlling neural crest delamination and migration as well as differentiation programs for neural crest derivatives like cartilage, pigment cells and peripheral neurons. B. Cross sections showing neurulation process in amphioxus embryos. In late gastrula, the embryo is ovoid with presumptive neural plate (green) becoming flattened. The boundaries of presumptive neural plate and epidermis (blue) at this stage are indicated by arrows, and the mesendoderm is colored in yellow. In early neurula, the epidermis (blue) at the edges of the neural plate detaches from the neural plate and begins to overgrow the neural plate, and the somites (pink) and notochord (red) start to form. Eventually the overgrowing dorsal epidermis fuse together at the dorsal midline, and the underneath neural plate curls up to form a dorsal, hollow neural tube in late neurula stage. Now, neurulation is complete, and the rudiment of notochord, anterior somites, as well as the endodermal tube rudiment that will form the pharyngeal tube and gut (yellow) become distinguishable.
2 Figure S2 Dose-dependent effect of zbmp4 treatment on amphioxus epidermal and neural gene expression. Whole-mount in situ hybridization for Dlx and SoxB1-a at late gastrula stage. All embryos are oriented in dorsal view with anterior at the top. Scale bar, 50 µm. Expression of epidermal marker Dlx was expanded by zbmp4 treatment in a dose-dependant manner; conversely, expression of neural marker SoxB1-a was reduced by low concentration of zbmp4 treatment and completely disappeared by high concentration of zbmp4.
3 Figure S3 Expression of Mitf, Tyrosinase, and two Tyrosinase-related proteins in amphioxus embryos. All embryos are oriented in dorsal view with anterior at the top. Scale bar, 50 µm. All these four genes are co-expressed throughout the epidermal ectoderm in gastrula stage and early neurula stage. During neurula stage, a patch of cells inside the developing neural tube (indicating by arrows) start to expressed these four genes, and the onset of Mitf transcription factor gene expression is earlier than the other melanogenic genes.
4 FGF8/17/18 subfamily Figure S4 Phylogenetic tree of FGF proteins constructed by MEGA program using the neighbor-joining method with default settings. Amphioxus protein is shown by large black dots. The number matching each branch indicates the percentage of times that a node was supported in 0 bootstrap pseudoreplications. Protein names are as explained in the Methods. The scale bar indicates an evolutionary distance of 0.2 amino acid substitutions per position. FGF8 has been implicated in playing roles in neural induction and neural crest development. Our survey identified an amphioxus candidate belonging to the FGF8/17/18 subfamily with high bootstrap value of %.
5 Q9NR61 DLL4 Hs O00548 DLL1 Hs Q9NYJ7 DLL3 Hs Delta estext fgenesh2 pg.c Delta Bf P41 delta Dm P18168 Serrate Dm 75 e gw Jagged Bf P78504 JAG1 Hs Q9Y219 JAG2 Hs Jagged/Serrate 0.1 Figure S5 Phylogenetic tree of Delta/Jagged/Serrate family proteins generated by MEGA program using the neighbor-joining method with default settings. Amphioxus proteins are shown by large black dots. The number matching each branch indicates the percentage of times that a node was supported in 0 bootstrap pseudoreplications. The unrooted tree is shown as a rooted tree for simplicity. Protein names are as explained in the Methods. The scale bar indicates an evolutionary distance of 0.1 amino acid substitutions per position. Delta and Jagged/Serrate proteins are ligands for Notch receptors. Our survey in amphioxus genome identified one candidate gene for Delta and one for Jagged/Serrate. The robust grouping of the tree supports their orthology assignment.
6 estext GenewiseH 1.C SoxB2 Bf AAY51532 SoxB2.3 Dm Q24533 SoxB2.1 Dm AAL13542 SoxB2.2 Dm Q9Y651 SOX21 Hs O95416 SOX14 Hs e gw SoxB1-c Bf Sox B group NP SoxNeuro Dm P41225 SOX3 Hs O00570 SOX1 Hs P48431 SOX2 Hs estext fgenesh2 kg.c SoxB1-a Bf estext fgenesh2 pg.c SoxB1-b Bf O60248 SOX15 Hs Sox G group Q05066 SRY Hs Sox A group 57 Q06945 SOX4 Hs O15370 SOX12 Hs Sox C group P35716 SOX11 Hs Q9UN79 SOX13 Hs 66 P35711 SOX5 Hs P35712 SOX6 Hs Sox D group Q9H6I2 SOX17 Hs Q9BT81 SOX7 Hs P35713 SOX18 Hs Sox F group 52 CAB63903 SoxE Dm P57073 SOX8 Hs P56693 SOX10 Hs P48436 SOX9 Hs Sox E group e gw SoxE Bf O94993 SOX30 Hs Sox H group 0.1 Figure S6 Phylogenetic tree of Sox proteins constructed by the neighbor-joining method with default settings. Amphioxus protein are shown by large black dots. The number matching each branch indicates the percentage of times that a node was supported in 0 bootstrap pseudoreplications. Protein names are as explained in the Methods. The scale bar indicates an evolutionary distance of 0.1 amino acid substitutions per position. Our survey of amphioxus Sox family genes was focused on the SoxB and SoxE groups. We identified an amphioxus SoxE group candidate which groups with vertebrate SoxE proteins with high bootstrap value(99%). We identified four candidates of amphioxus SoxB members. Although the bootstrap value of SoxB grouping was not very high, we designated the protein names based on the best hit analysis by blast search against a human proteome.
7 estext fgenesh2 pg.c Olig-c Bf estext fgenesh2 pg.c Olig-b Bf estext fgenesh2 pg.c Olig-a Bf Olig 94 Q7RTU3 OLIG3 Hs Q13516 OLIG2 Hs fgenesh2 pg.scaffold Beta3 Bf Q92886 Neurogenin1 Hs Q8NFJ8 BHLH5 Hs Q9Y4Z2 Neurogenin3 Hs Q9H2A3 Neurogenin2 Hs Q8C6A8 BETA3 Mm Neurogenin Beta3 estext fgenesh2 pg.c Neurogenin Bf 66 gw NeuroD Bf 99 Q9HD90 NDF4 Hs Q13562 NDF1 Hs NeuroD 97 Q15784 NDF2 Hs Q9JJR7 ASCL3 Mm P50553 ASCL1 Hs Q99929 ASCL2 Hs P09775 Asense Dm e gw Ash-a-like1 Bf e gw Ash-a-like2 Bf Achaete-Scute (Ash) P83 Achaete Dm P09774 Lsc Dm P84 Scute Dm 0.1 Figure S7 Phylogenetic tree of Proneural bhlh factors generated by the neighbor-joining method with default settings. Amphioxus proteins are shown by large black dots. The number matching each branch indicates the percentage of times that a node was supported in 0 bootstrap pseudo-replications. Protein names are as explained in the Methods. The scale bar indicates an evolutionary distance of 0.1 amino acid substitutions per position. We identified three possible Olig family orthologues in the amphioxus genome and one candidate for the closely related Beta3 family. The three amphioxus Olig genes are tandemly aligned in the genome within a 62-kb region, suggesting that these genes occurred by amphioxus-specific gene duplications. We also identified amphioxus orthologues of Neurogenin and NeuroD family with strong bootstrap values. There are two possible Ash family members in amphioxus genome, however, the phylogenetic resolution was low within the family. Therefore, we designated these two factors as Ash-a-like1 and Ash-a-like2 based on the best hit analysis.
8 82 P19484 TFEB Hs 66 O14948 TFECL Hs P19532 TFE3 Hs O75030 MITF Hs e gw Mitf Bf Mitf AAQ01726 Mitf Dm P22415 USF1 Hs fgenesh2 pm.scaffold USF Bf USF 65 Q15853 USF2 Hs 0.2 Figure S8 Phylogenetic tree of Mitf family proteins generated by MEGA program using the neighbor-joining method with default settings. Amphioxus proteins are shown by large black dots. The number matching each branch indicates the percentage of times that a node was supported in 0 bootstrap pseudoreplications. The unrooted tree is shown as a rooted tree for simplicity. Protein names are as explained in the Methods. The scale bar indicates an evolutionary distance of 0.2 amino acid substitutions per position. Our survey in amphioxus genome identified one orthologue of Mitf, and it was included with other Mitf family factors as a clade with high bootstrap support (%). We also identified a candidate for the USF family member, which is a closely related family to Mitf.
9 82 Q9W4S7 MYC Dm P01106 MYC Hs fgenesh2 pm.scaffold Myc Bf P04198 MYCN Hs P12524 MYCL1 Hs Myc P12525 MYCL2 Hs fgenesh2 pg.scaffold Max-a Bf gw Max-b Bf P61244 MAX Hs Max AAF49179 Max Dm 99 fgenesh2 pg.scaffold Mnt Bf Q99583 MNT Hs Mnt AAH00745 MAD3 Hs fgenesh2 pg.scaffold MAD Bf Q05195 MAD1 Hs Q14582 MAD4 Hs MAD 55 P50539 MXI1 Hs 0.2 Figure S9 Phylogenetic tree of Myc, Max, Mnt, and MAD family proteins generated by the neighbor-joining method with default settings. Amphioxus proteins are shown by large black dots. The number matching each branch indicates the percentage of times that a node was supported in 0 bootstrap pseudoreplications. The unrooted tree is shown as a rooted tree for simplicity. Protein names are as explained in the Methods. The scale bar indicates an evolutionary distance of 0.2 amino acid substitutions per position. Our survey in amphioxus genome identified one candidate orthologue of Myc family factor with good bootstrap support (%). We also identified orthologues of the closely related Max, Mnt, and MAD family members from amphioxus genome. There are two amphioxus Max members and the phylogenetic analysis suggests they occurred by amphioxus-specific duplication.
10 P40126 TYRP2 Hs 53 P29812 TYRP2 Mm P17643 TYRP1 Hs P07147 TYRP1 Mm BAC76423 HrTYR Hr Tyrp estext fgenesh2 pg.c Tyrp-b Bf fgenesh2 pg.scaffold Tyrp-a Bf BAC76424 HrTRP Hr 56 estext gwp.c Tyrosinase Bf P14679 TYRO Hs P11344 TYRO Mm Tyrosinase 0.1 Figure S10 Phylogenetic tree of Tyrosinase and Tyrosinase-related proteins (Tyrp) generated by the neighbor-joining method with default settings. Amphioxus proteins are shown by large black dots. The number matching each branch indicates the percentage of times that a node was supported in 0 bootstrap pseudoreplications. The unrooted tree is shown as a rooted tree for simplicity. Protein names are as explained in the Methods. The scale bar indicates an evolutionary distance of 0.1 amino acid substitutions per position. Tyrosinase and Tyrosinase-related proteins are separated clearly as two groups in our phylogenetic tree. Our phylogenetic analysis suggests that the tow Tyrosinase-related proteins we identified are amphioxus specific duplications, therefore we designated them as Tyrp-a and Tyrp-b.
11 83 P61586 RHOA Hs 69 P08134 RHOC Hs P48148 RHO1 Dm P62745 RHOB Hs fgenesh2 pg.scaffold RhoA/B/C-b RhoA/B/C estext fgenesh2 pm.c RhoA/B/C-a P84095 RHOG Hs fgenesh2 pm.scaffold RhoD/F Bf O00212 RHOD Hs Q9HBH0 RHOF Hs RhoD/F Q92730 RRHO6 Hs 87 P61587 RHOE Hs P52198 RHON Hs RhoE Q15669 RHOH Hs 0.1 Figure S11 Phylogenetic tree of Rho small G-proteins generated by MEGA program using the neighbor-joining method with default settings. Amphioxus proteins are shown by large black dots. The number matching each branch indicates the percentage of times that a node was supported in 0 bootstrap pseudoreplications. The unrooted tree is shown as a rooted tree for simplicity. Protein names are as explained in the Methods. The scale bar indicates an evolutionary distance of 0.1 amino acid substitutions per position. We identified two candidate gene models encoding Rho-like G-proteins belonging to the RhoA/B/C group (bootstrap value of 98%). There is also a possible RhoD/F orthologue in amphioxus, however, the bootstrap supporting value is not very high.
12 P21802 FGFR2 Hs P11362 FGFR1 Hs AAB33460 c-ret Gg AAG00544 c-ret Xl NP c-ret Dr P07949 RET Hs SAL fgenesh2 pg.scaffold Ret Bf AAF53977 Ret Dm XP LOC Am outgroup 0.1 Figure S12 Phylogenetic tree of Ret receptor tyrosine kinase proteins generated by the neighbor-joining method based on the alignment of a tyrosine kinase domain. Amphioxus protein is shown by the large black dot. The number matching each branch indicates the percentage of times that a node was supported in 0 bootstrap pseudoreplications. Protein names are as explained in the Methods. Human FGF receptors were used as outgroup proteins. The scale bar indicates an evolutionary distance of 0.1 amino acid substitutions per position. Our survey in amphioxus genome identified one candidate belonging to Ret family.
13 P21860 ERBB3 Hs Q15303 ERBB4 Hs P00533 EGFR Hs P04626 ERBB2 Hs EGF receptor estext fgenesh2 pg.c Erbb Bf 90 P04412 EGFR Dm P08069 IGF1R Hs P09208 INSR Dm outgroup 0.2 Figure S13 Phylogenetic tree of EGF receptors generated by the neighbor-joining method based on the alignment of a tyrosine kinase domain. Amphioxus protein is shown by large black dots. The number matching each branch indicates the percentage of times that a node was supported in 0 bootstrap pseudoreplications. Protein names are as explained in the Methods. Human and fly Insulin receptors were used as outgroup proteins. The scale bar indicates an evolutionary distance of 0.2 amino acid substitutions per position. In vertebrates, there are four members of epidermal growth factor (EGF) receptor family of receptor tyrosine kinase including ERBB1 (EGFR), ERBB2, ERBB3 and ERBB4. We identified only one amphioxus Erbb candidate in the genome, and the orthology among this amphioxus protein and EGFRs from other organisms is supported by high bootstrap value of %.
14 Table S1. Homologues of vertebrate neural crest network genes in the amphioxus Brachiosoma floridae genome Gene name The best hit model in the version The best hit protein in the human proteome cdna clone ID 1.0 assembly Neural plate border induction signals BMP2/4 estext_fgenesh2_pg.c_ P12643 BMP-2 bfga034p10 Wnt1 estext_fgenesh2_pm.c_ P04628 WNT-1 bfne123k05 Wnt3 estext_fgenesh2_pg.c_ P56704 WNT-3A bflv058m15 Wnt6 fgenesh2_pm.scaffold_ Q9Y6F9 WNT-6 bflv009a17 Wnt7 fgenesh2_kg.scaffold_00002 P56706 WNT-7B bfne126m18 Wnt8 fgenesh2_kg.scaffold_ Q93098 WNT-8B bfne137b06 FGF8/17/18 estext_fgenesh2_pg.c_ O60258 Fibroblast growth factor 17 bfne018g21 Notch estext_gwp.c_ Q5SXM3 NOTCH1 bfne093p03 Delta estext_fgenesh2_pg.c_ O00548 Delta-like 1 bfga003d05 Neural plate border specifiers Dlx estext_genewiseh_1.c_ P56177 DLX-1 bflv011k11 Msx estext_gwp.c_ P28360 MSX-1 bflv048l06 Zic (AmphiZic) estext_gwp.c_ Q2M3N1 ZIC1 bfne006l12 Pax3/7 fgenesh2_kg.scaffold_ Q2PJS5 PAX7 bfne062d03 Neural patterning/differentiation genes SoxB1-a * estext_fgenesh2_kg.c_ P48431 SOX-2 bfne037g13 SoxB1-b estext_fgenesh2_pg.c_ P48431 SOX-2 bfne072e07 SoxB1-c e_gw P48431 SOX-2 bfne126o15 SoxB2 estext_genewiseh_1.c_ O95416 SOX-14 bfad048c06 NeuroD gw Q13562 Neurogenic differentiation factor 1 N/A Neurogenin estext_fgenesh2_pg.c_90049 Q92886 Neurogenin 1 N/A Ash-a-like1 e_gw P50553 Achaete-scute homolog 1 N/A Ash-a-like2 e_gw P50553 Achaete-scute homolog 1 N/A Beta3 fgenesh2_pg.scaffold_ Q8NFJ8 Basic helix-loop-helix protein 5 N/A Olig-a estext_fgenesh2_pg.c_ Q7RTU3 Oligodendrocyte transcription factor 3 bfne042a07 Olig-b estext_fgenesh2_pg.c_ Q7RTU3 Oligodendrocyte transcription factor 3 bfne044b12 Olig-c estext_fgenesh2_pg.c_ Q7RTU3 Oligodendrocyte transcription factor 3 bfne004n21 Islet estext_fgenesh2_kg.c_ P61371 Islet-1 bfne013h18 Hu/elav fgenesh2_pg.scaffold_ Q12926 ELAV-like protein 2 bflv027g23
15 Neural crest specifiers Snail estext_fgenesh2_kg.c_ O43623 Zinc finger protein SLUG bfne095n03 FoxD estext_fgenesh2_kg.c_ Q9UJU5 FOXD3 bfne002k18 Twist fgenesh2_pm.scaffold_ Q8WVJ9 Twist-related protein 2 bfne115j15 Myc fgenesh2_pm.scaffold_ P01106 Myc proto-oncogene protein bfne011k03 Id fgenesh2_pg.scaffold_ Q02363 ID-2 bfne018j05 AP-2 e_gw P05549 AP-2 alpha bfne111f20 SoxE e_gw P48436 SOX-9 bfne111n21 Neural crest effector genes Mitf e_gw O75030 MITF bflv016p23 Tyrosinase estext_gwp.c_ P14679 Tyrosinase bfga039h23 Tyrp1 fgenesh2_pg.scaffold_ P17643 TRP-1 bfga030e07 Tyrp2 estext_fgenesh2_pg.c_ P40126 L-dopachrome tautomerase precursor bfga006j06 RhoA/B/C-a estext_fgenesh2_pm.c_ P61586 RHOA bfga002n08 RhoA/B/C-b fgenesh2_pg.scaffold_ P61586 RHOA bfne019m11 Ret SAL_fgenesh2_pg.scaffold_ P07949 RET N/A Erbb estext_fgenesh2_pg.c_ P21860 ErbB-3 N/A Col2a fgenesh2_pg.scaffold_53000 P02458 alpha 1 type II collagen bflv014e16 ckit Myelin protein P0 Not found Not found N/A: not available * This gene was named AmphiSox1/2/3 in previous literatures This gene model was annotated manually by combining two predicted gene models fgenesh2_pg.scaffold_ and fgenesh2_pg.scaffold_
16 Table S2. Expression of EGFP in chick embryos electroporated at stage 3+ to 4. Stage examined Somites/paraxial mesoderm Notochord Neural tube (hindbrain) Head mesenchyme Stage 8 (n=4) 4/4 4/4 0/4 4/4 Stage 10 (n=5) 5/5 3/5* 5/5 4/5** Stage 11 (n=4) 4/4 4/4 4/4 4/4 * The two embryos with no EGFP in the notochord did not show any expression of RFP in the notochord. ** The embryo with no EGFP expression in the head mesenchyme did not show any expression of RFP in the head mesenchyme.
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
9/4/2015 INDUCTION CHAPTER 1. Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology. Fig 1.
 INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
PRACTICE EXAM. 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos.
 PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 7, 2006 Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of BMPs more lateral mesoderm
Biology 4361 Paraxial and Intermediate Mesoderm December 7, 2006 Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of BMPs more lateral mesoderm
Bio 127 Section I Introduction to Developmental Biology. Cell Cell Communication in Development. Developmental Activities Coordinated in this Way
 Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
Cell Cell Communication in Development
 Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Role of Organizer Chages in Late Frog Embryos
 Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Sonic hedgehog (Shh) signalling in the rabbit embryo
 Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Developmental Biology 3230 Midterm Exam 1 March 2006
 Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning
 MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development June 23, 2009 Concepts Cell-Cell Communication Cells develop in the context of their environment, including: - their immediate
Biology 4361 - Developmental Biology Cell-Cell Communication in Development June 23, 2009 Concepts Cell-Cell Communication Cells develop in the context of their environment, including: - their immediate
Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio
 Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Supplementary Figures
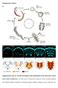 Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Name. Biology Developmental Biology Winter Quarter 2013 KEY. Midterm 3
 Name 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2013 KEY Midterm 3 Read the Following Instructions: * Answer 20 questions (5 points each) out of the available 25 questions
Name 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2013 KEY Midterm 3 Read the Following Instructions: * Answer 20 questions (5 points each) out of the available 25 questions
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome
 Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome Marcos S. Simões-Costa 1., Sonja J. McKeown 1., Joanne Tan-Cabugao 1, Tatjana Sauka-Spengler
Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome Marcos S. Simões-Costa 1., Sonja J. McKeown 1., Joanne Tan-Cabugao 1, Tatjana Sauka-Spengler
Question Set # 4 Answer Key 7.22 Nov. 2002
 Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm July 28, 2008 Paraxial and Intermediate Mesoderm Overview Development of major mesodermal lineages Somites: formation specification and differentiation Mesodermal
Biology 4361 Paraxial and Intermediate Mesoderm July 28, 2008 Paraxial and Intermediate Mesoderm Overview Development of major mesodermal lineages Somites: formation specification and differentiation Mesodermal
NEURAL CREST SPECIFICATION: MIGRATING INTO GENOMICS
 NEURAL CREST SPECIFICATION: MIGRATING INTO GENOMICS Laura S. Gammill and Marianne Bronner-Fraser The bones in your face, the pigment in your skin and the neural circuitry that controls your digestive tract
NEURAL CREST SPECIFICATION: MIGRATING INTO GENOMICS Laura S. Gammill and Marianne Bronner-Fraser The bones in your face, the pigment in your skin and the neural circuitry that controls your digestive tract
Developmental processes Differential gene expression Introduction to determination The model organisms used to study developmental processes
 Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Bio Section III Organogenesis. The Neural Crest and Axonal Specification. Student Learning Objectives. Student Learning Objectives
 Bio 127 - Section III Organogenesis The Neural Crest and Axonal Specification Gilbert 9e Chapter 10 Student Learning Objectives 1. You should understand that the neural crest is an evolutionary advancement
Bio 127 - Section III Organogenesis The Neural Crest and Axonal Specification Gilbert 9e Chapter 10 Student Learning Objectives 1. You should understand that the neural crest is an evolutionary advancement
Life Sciences For NET & SLET Exams Of UGC-CSIR. Section B and C. Volume-08. Contents A. BASIC CONCEPT OF DEVELOPMENT 1
 Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Lecture 3 - Molecular Regulation of Development. Growth factor signaling, Hox genes and the body plan
 Lecture 3 - Molecular Regulation of Development. Growth factor signaling, Hox genes and the body plan Lecture Objectives Outline August 18, 2015, M.D., Ph.D. To understand how cell differentiation and
Lecture 3 - Molecular Regulation of Development. Growth factor signaling, Hox genes and the body plan Lecture Objectives Outline August 18, 2015, M.D., Ph.D. To understand how cell differentiation and
Name KEY. Biology Developmental Biology Winter Quarter Midterm 3 KEY
 Name KEY 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2009 Midterm 3 KEY All of the 25 multi-choice questions are single-answer. Choose the best answer. (4 pts each) Place
Name KEY 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2009 Midterm 3 KEY All of the 25 multi-choice questions are single-answer. Choose the best answer. (4 pts each) Place
UNIVERSITY OF YORK BIOLOGY. Developmental Biology
 Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
Skeletal Development in Human
 Atlas of Genetics and Cytogenetics in Oncology and Haematology Skeletal Development in Human Skeletal development in human - Long version I. Introduction I.1 Developmental genes in Drosophila I.2 Skeletal
Atlas of Genetics and Cytogenetics in Oncology and Haematology Skeletal Development in Human Skeletal development in human - Long version I. Introduction I.1 Developmental genes in Drosophila I.2 Skeletal
Chapter 2 Embryological Origins and the Identification of Neural Crest Cells
 Chapter 2 Embryological Origins and the Identification of Neural Crest Cells Neural Crest This chapter seeks to answer a twofold question: When and by what mechanisms do the neural crest (NC) and neural
Chapter 2 Embryological Origins and the Identification of Neural Crest Cells Neural Crest This chapter seeks to answer a twofold question: When and by what mechanisms do the neural crest (NC) and neural
!!!!!!!! DB3230 Midterm 2 12/13/2013 Name:
 1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
Conclusions. The experimental studies presented in this thesis provide the first molecular insights
 C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
1. What are the three general areas of the developing vertebrate limb? 2. What embryonic regions contribute to the developing limb bud?
 Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Neural development its all connected
 Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8
 Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Neural Crest Development. Prof. Ken Ashwell Department of Anatomy, School of Medical Sciences
 Neural Crest Development Prof. Ken Ashwell Department of Anatomy, School of Medical Sciences Lecture plan What is the neural crest? Where does neural crest come from? DerivaBves of the neural crest MigraBon
Neural Crest Development Prof. Ken Ashwell Department of Anatomy, School of Medical Sciences Lecture plan What is the neural crest? Where does neural crest come from? DerivaBves of the neural crest MigraBon
Exam 3 (Final Exam) December 20, 2007
 Biology 4361 Exam 3 (Final Exam) December 20, 2007 Name: ID: Multiple choice (1 point each. Indicate the best answer.) 1. During Drosophila gastrulation, mesoderm moves in through the a. primitives streak.
Biology 4361 Exam 3 (Final Exam) December 20, 2007 Name: ID: Multiple choice (1 point each. Indicate the best answer.) 1. During Drosophila gastrulation, mesoderm moves in through the a. primitives streak.
NIH Public Access Author Manuscript Histochem Cell Biol. Author manuscript; available in PMC 2013 August 01.
 NIH Public Access Author Manuscript Published in final edited form as: Histochem Cell Biol. 2012 August ; 138(2): 179 186. doi:10.1007/s00418-012-0999-z. Formation and migration of neural crest cells in
NIH Public Access Author Manuscript Published in final edited form as: Histochem Cell Biol. 2012 August ; 138(2): 179 186. doi:10.1007/s00418-012-0999-z. Formation and migration of neural crest cells in
A simple molecular model of neurulation
 A simple molecular model of neurulation Michel Kerszberg* and Jean-Pierre Changeux Summary A molecular model for the morphogenesis of the central nervous system is built and solved by computer. The formalism
A simple molecular model of neurulation Michel Kerszberg* and Jean-Pierre Changeux Summary A molecular model for the morphogenesis of the central nervous system is built and solved by computer. The formalism
4. Neural tube cells are specified by opposing dorsal-ventral gradients of a. Wnts and Nodal. b. FGF and Shh. c. BMPs and Wnts. d. BMPs and Shh.
 Biology 4361 Name: KEY Exam 4 ID#: August 1, 2008 Multiple choice (one point each; indicate the best answer) 1. Neural tube closure is accomplished by movement of the a. medial hinge point cells. b. medial
Biology 4361 Name: KEY Exam 4 ID#: August 1, 2008 Multiple choice (one point each; indicate the best answer) 1. Neural tube closure is accomplished by movement of the a. medial hinge point cells. b. medial
THE SPECIFICATION OF DORSAL CELL FATES IN THE VERTEBRATE CENTRAL NERVOUS SYSTEM
 Annu. Rev. Neurosci. 1999. 22:261 94 Copyright c 1999 by Annual Reviews. All rights reserved THE SPECIFICATION OF DORSAL CELL FATES IN THE VERTEBRATE CENTRAL NERVOUS SYSTEM Kevin J. Lee and Thomas M. Jessell
Annu. Rev. Neurosci. 1999. 22:261 94 Copyright c 1999 by Annual Reviews. All rights reserved THE SPECIFICATION OF DORSAL CELL FATES IN THE VERTEBRATE CENTRAL NERVOUS SYSTEM Kevin J. Lee and Thomas M. Jessell
Research article. Summary. Introduction. Kaoru S. Imai*, Kyosuke Hino*, Kasumi Yagi, Nori Satoh and Yutaka Satou
 Research article 4047 Gene expression profiles of transcription factors and signaling molecules in the ascidian embryo: towards a comprehensive understanding of gene networks Kaoru S. Imai*, Kyosuke Hino*,
Research article 4047 Gene expression profiles of transcription factors and signaling molecules in the ascidian embryo: towards a comprehensive understanding of gene networks Kaoru S. Imai*, Kyosuke Hino*,
Maternal Control of GermLayer Formation in Xenopus
 Maternal Control of GermLayer Formation in Xenopus The zygotic genome is activated at the mid-blastula transition mid-blastula fertilized egg Xenopus gastrulae early-gastrula 7 hrs 10 hrs control not VP
Maternal Control of GermLayer Formation in Xenopus The zygotic genome is activated at the mid-blastula transition mid-blastula fertilized egg Xenopus gastrulae early-gastrula 7 hrs 10 hrs control not VP
Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis
 Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis 10 December 2012 - Corrections - Exercise 1 Non-vertebrate chordates generally possess 2 homologs, vertebrates 3 or more gene copies; a Drosophila
Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis 10 December 2012 - Corrections - Exercise 1 Non-vertebrate chordates generally possess 2 homologs, vertebrates 3 or more gene copies; a Drosophila
Biology 218, practise Exam 2, 2011
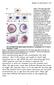 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Elucidation of the Evolutionary Origin of the Neural Crest. William K. Nesbitt
 Elucidation of the Evolutionary Origin of the Neural Crest William K. Nesbitt Under the supervision of Dr. David McClay, Department of Biology, Duke University April 22, 2016 Research Supervisor Faculty
Elucidation of the Evolutionary Origin of the Neural Crest William K. Nesbitt Under the supervision of Dr. David McClay, Department of Biology, Duke University April 22, 2016 Research Supervisor Faculty
Developmental Biology Lecture Outlines
 Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Figure S1: Phylogenetic analysis of the Platynereis Notch pathway core components, Pdu-
 Electronic supplementary material legends from The Notch pathway in the annelid Platynereis: Insights into chaetogenesis and neurogenesis processes ; Eve Gazave, Quentin I. B. Lemaître and Guillaume Balavoine;
Electronic supplementary material legends from The Notch pathway in the annelid Platynereis: Insights into chaetogenesis and neurogenesis processes ; Eve Gazave, Quentin I. B. Lemaître and Guillaume Balavoine;
Neural crest development is regulated by the transcription factor Sox9
 Research article 5681 Neural crest development is regulated by the transcription factor Sox9 Martin Cheung and James Briscoe* Developmental Neurobiology, National Institute for Medical Research, Mill Hill,
Research article 5681 Neural crest development is regulated by the transcription factor Sox9 Martin Cheung and James Briscoe* Developmental Neurobiology, National Institute for Medical Research, Mill Hill,
Tsukushi Modulates Xnr2, FGF and BMP Signaling: Regulation of Xenopus Germ Layer Formation
 Tsukushi Modulates Xnr2, FGF and BMP Signaling: Regulation of Xenopus Germ Layer Formation Samantha A. Morris 1 *, Alexandra D. Almeida 1, Hideaki Tanaka 2, Kunimasa Ohta 2, Shin-ichi Ohnuma 1 * 1 Department
Tsukushi Modulates Xnr2, FGF and BMP Signaling: Regulation of Xenopus Germ Layer Formation Samantha A. Morris 1 *, Alexandra D. Almeida 1, Hideaki Tanaka 2, Kunimasa Ohta 2, Shin-ichi Ohnuma 1 * 1 Department
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 Evolution and development ("evo-devo") The last frontier in our understanding of biological forms is an understanding
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 Evolution and development ("evo-devo") The last frontier in our understanding of biological forms is an understanding
Early Development in Invertebrates
 Developmental Biology Biology 4361 Early Development in Invertebrates October 25, 2006 Early Development Overview Cleavage rapid cell divisions divisions of fertilized egg into many cells Gastrulation
Developmental Biology Biology 4361 Early Development in Invertebrates October 25, 2006 Early Development Overview Cleavage rapid cell divisions divisions of fertilized egg into many cells Gastrulation
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A)
 Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
craniofacial development and WNT signaling. Shachi Bhatt
 MED23: a Mediator subunit's role in global gene transcription, regulation of craniofacial development and WNT signaling. BY 2013 Shachi Bhatt Submitted to the graduate degree program in Anatomy and Cell
MED23: a Mediator subunit's role in global gene transcription, regulation of craniofacial development and WNT signaling. BY 2013 Shachi Bhatt Submitted to the graduate degree program in Anatomy and Cell
Bioinformatics tools for phylogeny and visualization. Yanbin Yin
 Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Spatiotemporal regulation of nervous system development in the annelid Capitella teleta
 DOI 10.1186/s13227-017-0076-8 EvoDevo RESEARCH Spatiotemporal reguion of nervous system development in the annelid Capitella teleta Abhinav Sur 1, Craig R. Magie 2, Elaine C. Seaver 3 and Néva P. Meyer
DOI 10.1186/s13227-017-0076-8 EvoDevo RESEARCH Spatiotemporal reguion of nervous system development in the annelid Capitella teleta Abhinav Sur 1, Craig R. Magie 2, Elaine C. Seaver 3 and Néva P. Meyer
Roles of retinoic acid and Tbx1/10 in pharyngeal segmentation: amphioxus and the ancestral chordate condition
 Koop et al. EvoDevo 2014, 5:36 RESEARCH Open Access Roles of retinoic acid and Tbx1/10 in pharyngeal segmentation: amphioxus and the ancestral chordate condition Demian Koop 1, Jie Chen 2, Maria Theodosiou
Koop et al. EvoDevo 2014, 5:36 RESEARCH Open Access Roles of retinoic acid and Tbx1/10 in pharyngeal segmentation: amphioxus and the ancestral chordate condition Demian Koop 1, Jie Chen 2, Maria Theodosiou
presumptiv e germ layers during Gastrulatio n and neurulation Somites
 Vertebrate embryos are similar at the phylotypic stage Patterning the Vertebrate Body Plan II: Mesoderm & Early Nervous System Wolpert L, Beddington R, Jessell T, Lawrence P, Meyerowitz E, Smith J. (2001)
Vertebrate embryos are similar at the phylotypic stage Patterning the Vertebrate Body Plan II: Mesoderm & Early Nervous System Wolpert L, Beddington R, Jessell T, Lawrence P, Meyerowitz E, Smith J. (2001)
Exam 2 ID#: November 9, 2006
 Biology 4361 Name: KEY Exam 2 ID#: November 9, 2006 Multiple choice (one point each) Circle the best answer. 1. Inducers of Xenopus lens and optic vesicle include a. pharyngeal endoderm and anterior neural
Biology 4361 Name: KEY Exam 2 ID#: November 9, 2006 Multiple choice (one point each) Circle the best answer. 1. Inducers of Xenopus lens and optic vesicle include a. pharyngeal endoderm and anterior neural
Exam 4 ID#: July 7, 2008
 Biology 4361 Name: KEY Exam 4 ID#: July 7, 2008 Multiple choice (one point each; indicate the best answer) 1. RNA polymerase II is not able to transcribe RNA unless a. it is first bound to TFIIB. b. its
Biology 4361 Name: KEY Exam 4 ID#: July 7, 2008 Multiple choice (one point each; indicate the best answer) 1. RNA polymerase II is not able to transcribe RNA unless a. it is first bound to TFIIB. b. its
2/23/09. Regional differentiation of mesoderm. Morphological changes at early postgastrulation. Segments organize the body plan during embryogenesis
 Regional differentiation of mesoderm Axial Paraxial Intermediate Somatic Splanchnic Chick embryo Morphological changes at early postgastrulation stages Segments organize the body plan during embryogenesis
Regional differentiation of mesoderm Axial Paraxial Intermediate Somatic Splanchnic Chick embryo Morphological changes at early postgastrulation stages Segments organize the body plan during embryogenesis
Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural crest to definitive roof plate
 Washington University School of Medicine Digital Commons@Becker Open Access Publications 2016 Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural crest
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2016 Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural crest
THE PROBLEMS OF DEVELOPMENT. Cell differentiation. Cell determination
 We emphasize these points from Kandel in Bi/CNS 150 Bi/CNS/NB 150: Neuroscience Read Lecture Lecture Friday, October 2, 2015 Development 1: pp 5-10 Introduction Brains evolved All higher animals have brains
We emphasize these points from Kandel in Bi/CNS 150 Bi/CNS/NB 150: Neuroscience Read Lecture Lecture Friday, October 2, 2015 Development 1: pp 5-10 Introduction Brains evolved All higher animals have brains
10/2/2015. Chapter 4. Determination and Differentiation. Neuroanatomical Diversity
 Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Questions in developmental biology. Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration
 Questions in developmental biology Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration Representative cell types of a vertebrate zygote => embryo => adult differentiation
Questions in developmental biology Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration Representative cell types of a vertebrate zygote => embryo => adult differentiation
The mouse Ovol2 gene is required for cranial neural tube development
 Developmental Biology 291 (2006) 38 52 www.elsevier.com/locate/ydbio The mouse Ovol2 gene is required for cranial neural tube development Douglas R. Mackay a, Ming Hu a, Baoan Li a, Catherine Rhéaume a,
Developmental Biology 291 (2006) 38 52 www.elsevier.com/locate/ydbio The mouse Ovol2 gene is required for cranial neural tube development Douglas R. Mackay a, Ming Hu a, Baoan Li a, Catherine Rhéaume a,
10/15/09. Tetrapod Limb Development & Pattern Formation. Developing limb region is an example of a morphogenetic field
 Tetrapod Limb Development & Pattern Formation Figure 16.5(1) Limb Bud Formation derived from lateral plate (somatic) & paraxial (myotome) Fig. 16.2 Prospective Forelimb Field of Salamander Ambystoma maculatum
Tetrapod Limb Development & Pattern Formation Figure 16.5(1) Limb Bud Formation derived from lateral plate (somatic) & paraxial (myotome) Fig. 16.2 Prospective Forelimb Field of Salamander Ambystoma maculatum
A New Mechanistic Scenario for the Origin and Evolution of Vertebrate Cartilage
 A New Mechanistic Scenario for the Origin and Evolution of Vertebrate Cartilage Maria Cattell 1, Su Lai 1, Robert Cerny 2, Daniel Meulemans Medeiros 1 * 1 Department of Ecology and Evolutionary Biology,
A New Mechanistic Scenario for the Origin and Evolution of Vertebrate Cartilage Maria Cattell 1, Su Lai 1, Robert Cerny 2, Daniel Meulemans Medeiros 1 * 1 Department of Ecology and Evolutionary Biology,
Dorsoventral Patterning in Hemichordates: Insights into Early Chordate Evolution
 : Insights into Early Chordate Evolution The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters. Citation Published Version Accessed
: Insights into Early Chordate Evolution The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters. Citation Published Version Accessed
Developmental Biology
 Developmental Biology 340 (2010) 75 87 Contents lists available at ScienceDirect Developmental Biology journal homepage: www.elsevier.com/developmentalbiology A divergent Tbx6-related gene and Tbx6 are
Developmental Biology 340 (2010) 75 87 Contents lists available at ScienceDirect Developmental Biology journal homepage: www.elsevier.com/developmentalbiology A divergent Tbx6-related gene and Tbx6 are
DOWNLOAD PDF DETERMINATION OF PREPLACODAL ECTODERM AND SENSORY PLACODES SALLY A. MOODY
 Chapter 1 : Olfactory epithelium - Wikipedia The sensory organs of the vertebrate head derive from two embryological structures, the neural crest and the ectodermal placodes. Although quite a lot is known
Chapter 1 : Olfactory epithelium - Wikipedia The sensory organs of the vertebrate head derive from two embryological structures, the neural crest and the ectodermal placodes. Although quite a lot is known
Neural crest induction by paraxial mesoderm in Xenopus embryos requires FGF signals
 Development 130, 3111-3124 2003 The Company of Biologists Ltd doi:10.1242/dev.00531 3111 Neural crest induction by paraxial mesoderm in Xenopus embryos requires FGF signals Anne-Hélène Monsoro-Burq*, Russell
Development 130, 3111-3124 2003 The Company of Biologists Ltd doi:10.1242/dev.00531 3111 Neural crest induction by paraxial mesoderm in Xenopus embryos requires FGF signals Anne-Hélène Monsoro-Burq*, Russell
Orthology Part I: concepts and implications Toni Gabaldón Centre for Genomic Regulation (CRG), Barcelona
 Orthology Part I: concepts and implications Toni Gabaldón Centre for Genomic Regulation (CRG), Barcelona (tgabaldon@crg.es) http://gabaldonlab.crg.es Homology the same organ in different animals under
Orthology Part I: concepts and implications Toni Gabaldón Centre for Genomic Regulation (CRG), Barcelona (tgabaldon@crg.es) http://gabaldonlab.crg.es Homology the same organ in different animals under
A gene catalogue of the amphioxus nervous system
 Int. J. Biol. Sci. 2006, 2 149 Review A gene catalogue of the amphioxus nervous system Èlia Benito-Gutiérrez International Journal of Biological Sciences ISSN 1449-2288 www.biolsci.org 2006 2(3):149-160
Int. J. Biol. Sci. 2006, 2 149 Review A gene catalogue of the amphioxus nervous system Èlia Benito-Gutiérrez International Journal of Biological Sciences ISSN 1449-2288 www.biolsci.org 2006 2(3):149-160
The Radiata-Bilateria split. Second branching in the evolutionary tree
 The Radiata-Bilateria split Second branching in the evolutionary tree Two very important characteristics are used to distinguish between the second bifurcation of metazoans Body symmetry Germinal layers
The Radiata-Bilateria split Second branching in the evolutionary tree Two very important characteristics are used to distinguish between the second bifurcation of metazoans Body symmetry Germinal layers
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018
 Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Nature Genetics: doi: /ng Supplementary Figure 1. Icm/Dot secretion system region I in 41 Legionella species.
 Supplementary Figure 1 Icm/Dot secretion system region I in 41 Legionella species. Homologs of the effector-coding gene lega15 (orange) were found within Icm/Dot region I in 13 Legionella species. In four
Supplementary Figure 1 Icm/Dot secretion system region I in 41 Legionella species. Homologs of the effector-coding gene lega15 (orange) were found within Icm/Dot region I in 13 Legionella species. In four
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature12414 Supplementary Figure 1 a b Illumina 521M reads, single-end/76 bp Illumina 354M reads, paired-end/ bp raw read processing Trinity assembly primary assembly: 189,820 transcripts Frequency
doi:10.1038/nature12414 Supplementary Figure 1 a b Illumina 521M reads, single-end/76 bp Illumina 354M reads, paired-end/ bp raw read processing Trinity assembly primary assembly: 189,820 transcripts Frequency
Overexpression of Snail family members highlights their ability to promote chick neural crest formation
 Development 129, 1583-1593 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV1802 1583 Overexpression of Snail family members highlights their ability to promote chick neural crest
Development 129, 1583-1593 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV1802 1583 Overexpression of Snail family members highlights their ability to promote chick neural crest
Exam 3 ID#: July 31, 2009
 Biology 4361 Name: KEY Exam 3 ID#: July 31, 2009 Multiple choice (one point each; indicate the best answer) 1. Neural tube closure is accomplished by movement of the a. medial hinge point cells. b. medial
Biology 4361 Name: KEY Exam 3 ID#: July 31, 2009 Multiple choice (one point each; indicate the best answer) 1. Neural tube closure is accomplished by movement of the a. medial hinge point cells. b. medial
TCF/Lef regulates the Gsx ParaHox gene in central nervous system development in chordates
 Garstang et al. BMC Evolutionary Biology (2016) 16:57 DOI 10.1186/s12862-016-0614-3 RESEARCH ARTICLE Open Access TCF/Lef regulates the Gsx ParaHox gene in central nervous system development in chordates
Garstang et al. BMC Evolutionary Biology (2016) 16:57 DOI 10.1186/s12862-016-0614-3 RESEARCH ARTICLE Open Access TCF/Lef regulates the Gsx ParaHox gene in central nervous system development in chordates
Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural crest to definitive roof plate
 Nitzan et al. BMC Biology (2016) 14:23 DOI 10.1186/s12915-016-0245-6 RESEARCH ARTICLE Open Access Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural
Nitzan et al. BMC Biology (2016) 14:23 DOI 10.1186/s12915-016-0245-6 RESEARCH ARTICLE Open Access Dynamics of BMP and Hes1/Hairy1 signaling in the dorsal neural tube underlies the transition from neural
Unicellular: Cells change function in response to a temporal plan, such as the cell cycle.
 Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
Mesoderm Development
 Quiz rules: Spread out across available tables No phones, text books, or (lecture) notes on your desks No consultation with your colleagues No websites open other than the Quiz page No screen snap shots
Quiz rules: Spread out across available tables No phones, text books, or (lecture) notes on your desks No consultation with your colleagues No websites open other than the Quiz page No screen snap shots
Developmental Zoology. Ectodermal derivatives (ZOO ) Developmental Stages. Developmental Stages
 Developmental Zoology (ZOO 228.1.0) Ectodermal derivatives 1 Developmental Stages Ø Early Development Fertilization Cleavage Gastrulation Neurulation Ø Later Development Organogenesis Larval molts Metamorphosis
Developmental Zoology (ZOO 228.1.0) Ectodermal derivatives 1 Developmental Stages Ø Early Development Fertilization Cleavage Gastrulation Neurulation Ø Later Development Organogenesis Larval molts Metamorphosis
HHS Public Access Author manuscript Science. Author manuscript; available in PMC 2016 December 24.
 Reprogramming of avian neural crest axial identity and cell fate Marcos Simoes-Costa 1 and Marianne E. Bronner 1 1 Division of Biology and Biological Engineering, California Institute of Technology, Pasadena,
Reprogramming of avian neural crest axial identity and cell fate Marcos Simoes-Costa 1 and Marianne E. Bronner 1 1 Division of Biology and Biological Engineering, California Institute of Technology, Pasadena,
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Microarray Identification of Novel Downstream Targets of FoxD4L1/D5, a Critical Component of the Neural Ectodermal Transcriptional Network
 a DEVELOPMENTAL DYNAMICS 239:3467 3480, 2010 PATTERNS & PHENOTYPES Microarray Identification of Novel Downstream Targets of FoxD4L1/D5, a Critical Component of the Neural Ectodermal Transcriptional Network
a DEVELOPMENTAL DYNAMICS 239:3467 3480, 2010 PATTERNS & PHENOTYPES Microarray Identification of Novel Downstream Targets of FoxD4L1/D5, a Critical Component of the Neural Ectodermal Transcriptional Network
Origin of mesenchymal stem cell
 Origin of mesenchymal stem cell Takumi Era Department of Cell Modulation, Institute of Molecular embryology and Genetics (IMEG) Kumamoto University Differentiation pathways of in vitro ES cell culture
Origin of mesenchymal stem cell Takumi Era Department of Cell Modulation, Institute of Molecular embryology and Genetics (IMEG) Kumamoto University Differentiation pathways of in vitro ES cell culture
Developmental Biology Biology Ectodermal Organs. November 22, 2005
 Developmental Biology Biology 4361 Ectodermal Organs November 22, 2005 Germinal neuroepithelium external limiting membrane neural tube neuroepithelium (stem cells) Figure 13.3 Figure 13.4 Neuroepithelial
Developmental Biology Biology 4361 Ectodermal Organs November 22, 2005 Germinal neuroepithelium external limiting membrane neural tube neuroepithelium (stem cells) Figure 13.3 Figure 13.4 Neuroepithelial
Development of somites and their derivatives in amphioxus, and implications for the evolution of vertebrate somites
 Mansfield et al. EvoDevo (2015) 6:21 DOI 10.1186/s13227-015-0007-5 RESEARCH Open Access Development of somites and their derivatives in amphioxus, and implications for the evolution of vertebrate somites
Mansfield et al. EvoDevo (2015) 6:21 DOI 10.1186/s13227-015-0007-5 RESEARCH Open Access Development of somites and their derivatives in amphioxus, and implications for the evolution of vertebrate somites
Zebrafish foxd3 is selectively required for neural crest specification, migration and survival
 Developmental Biology 292 (2006) 174 188 www.elsevier.com/locate/ydbio Zebrafish foxd3 is selectively required for neural crest specification, migration and survival Rodney A. Stewart a, Brigitte L. Arduini
Developmental Biology 292 (2006) 174 188 www.elsevier.com/locate/ydbio Zebrafish foxd3 is selectively required for neural crest specification, migration and survival Rodney A. Stewart a, Brigitte L. Arduini
Ankyrin domain encoding genes from an ancient horizontal transfer are functionally integrated into Nasonia developmental gene regulatory networks
 Pers and Lynch Genome Biology (2018) 19:148 https://doi.org/10.1186/s13059-018-1526-x RESEARCH Open Access Ankyrin domain encoding genes from an ancient horizontal transfer are functionally integrated
Pers and Lynch Genome Biology (2018) 19:148 https://doi.org/10.1186/s13059-018-1526-x RESEARCH Open Access Ankyrin domain encoding genes from an ancient horizontal transfer are functionally integrated
Coordination of Cell Differentiation and Migration in Mathematical Models of Caudal Embryonic Axis Extension
 Coordination of Cell Differentiation and Migration in Mathematical Models of Caudal Embryonic Axis Extension Nigel C. Harrison 1, Ruth Diez del Corral 2 *, Bakhtier Vasiev 1 * 1 Department of Mathematical
Coordination of Cell Differentiation and Migration in Mathematical Models of Caudal Embryonic Axis Extension Nigel C. Harrison 1, Ruth Diez del Corral 2 *, Bakhtier Vasiev 1 * 1 Department of Mathematical
BLAST. Varieties of BLAST
 BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
Gene Duplication of the Zebrafish kit ligand and Partitioning of Melanocyte Development Functions to kit ligand a
 Gene Duplication of the Zebrafish kit ligand and Partitioning of Melanocyte Development Functions to kit ligand a Keith A. Hultman 1, Nathan Bahary 2, Leonard I. Zon 3,4, Stephen L. Johnson 1* 1 Department
Gene Duplication of the Zebrafish kit ligand and Partitioning of Melanocyte Development Functions to kit ligand a Keith A. Hultman 1, Nathan Bahary 2, Leonard I. Zon 3,4, Stephen L. Johnson 1* 1 Department
A Requirement for Retinoic Acid-Mediated Transcriptional Activation in Ventral Neural Patterning and Motor Neuron Specification
 Neuron, Vol. 40, 81 95, September 25, 2003, Copyright 2003 by Cell Press A Requirement for Retinoic Acid-Mediated Transcriptional Activation in Ventral Neural Patterning and Motor Neuron Specification
Neuron, Vol. 40, 81 95, September 25, 2003, Copyright 2003 by Cell Press A Requirement for Retinoic Acid-Mediated Transcriptional Activation in Ventral Neural Patterning and Motor Neuron Specification
