SUPPLEMENTAL DATA - 1. This file contains: Supplemental methods. Supplemental results. Supplemental tables S1 and S2. Supplemental figures S1 to S4
|
|
|
- Lily Boyd
- 5 years ago
- Views:
Transcription
1 Protein Disulfide Isomerase is Required for Platelet-Derived Growth Factor-Induced Vascular Smooth Muscle Cell Migration, Nox1 Expression and RhoGTPase Activation Luciana A. Pescatore 1, Diego Bonatto 2, Fábio L. Forti 3, Amine Sadok 4, Hervé Kovacic 4, Francisco R.M. Laurindo 1 1 Vascular Biology Laboratory, Heart Institute (InCor), University of São Paulo School of Medicine, São Paulo, Brazil, , 2 Biotechnology Center, Molecular and Cellular Biology Department, Federal University of Rio Grande do Sul, Brazil, 15005, 3 Chemistry Institute, University of São Paulo, Brazil, , 4 INSERM UMR911, Université de la Mediterranée, Marseille, France, Running title: PDI requirement for redox cell migration and GTPase activity To whom correspondence should be addressed: Francisco R. M. Laurindo, Heart Institute (InCor), University of São Paulo School of Medicine, Vascular Biology Laboratory. Av. Eneas Carvalho Aguiar, 44, Annex II, 9 th floor, CEP , São Paulo, Brazil. Phone: 55 (11) ; Fax: 55(11) ; francisco.laurindo@incor.usp.br SUPPLEMENTAL DATA - 1 This file contains: Supplemental methods Supplemental results Supplemental tables S1 and S2 Supplemental figures S1 to S4 1
2 SUPPLEMENTAL METHODS Physical protein-protein (PPPI) network design and global topological analysis The interactomic data gathered from human PDI proteins was used to obtain informations about their potential interactions with others proteins in the context of physical protein-protein interactions (PPPI networks) in Homo sapiens. In this sense, the data mining screening and network design of a major PPPI network (Figure S2) was performed using Cytoscape software, version (1). For this purpose, we used the PPPI data of H. sapiens available in STRING 9 ( database using the following parameters: active prediction methods all enabled except text mining; no more than 50 interactions; medium or standard confidence score (0.700 or 0.400, respectively); and network depth equal to 1 with addition of new nodes until saturation of the network. The PDI-associated PPPI networks obtained from this first screening were then combined in a unique PPPI network by employing the union function of the Cytoscape core plugin Merge Networks (Figure S2). The major PPPI network was then analyzed with Molecular Complex Detection (MCODE) software (2), a Cytoscape plugin available at in order to detect subnetworks or cluster of proteins that could represent distinct biologic processes. The parameters used for MCODE to generate the subnetworks were: loops included; degree cutoff 2; deletion of single connected nodes from cluster (haircut option enable); expansion of cluster by one neighbor shell allowed (fluff option enable); node density cutoff 0.1; node score cutoff 0.2; kcore 2; and maximum depth of network 100. Network centralities and local topological analyses A major network centrality (bottleneck nodes) was computed from the major PPPI networks using the Cytoscape plugin CytoHubba (3). A PPPI subnetwork containing the 50 major nodes with the highest bottleneck scores was drawn using the Cytoscape plugin Cyto-Hubba (3) and is available at Gene ontology analysis Gene ontology (GO) clustering analysis was performed using Biological Network Gene Ontology (BiNGO) (4) software, a Cytoscape plugin available at 2
3 The degree of functional enrichment for a given cluster and category was quantitatively assessed (p value) by hypergeometric distribution (5) and a multiple test correction was applied using the false discovery rate (FDR) (6) algorithm, fully implemented in BiNGO software. Overrepresented biological process categories were generated after FDR correction, with a significance level of SUPPLEMENTAL RESULTS The interactomic data about human PDI proteins obtained from different databases prompt us to ask how PDIs interacts with different proteins associated to oxidative stress. In this sense, a search for potential proteins and/or mechanisms and their associated biological processes that are affected by PDIs was initiated. To achieve this goal, different PPPI networks using Homo sapiens data were retrieved from STRING database. Shared proteins and subnetworks present in the major PPPI network (Figure S2) were identified and retrieved using the Cytoscape-associated plugin MCODE and subjected to a Gene Ontology (GO) analysis in order to obtain information about the nature and number of subgraphs belonging to the network and their associated biological processes. Results obtained from MCODE and GO analysis show that the final PPPI network (Figure S2) contains 159 nodes and 565 connectors and is composed by two interconnected clusters, each comprising different biological processes (Figures S3 and S4). GO analyses showed that these biological processes can be classified for cluster 1 into: (i) intracellular signal transduction, (ii) reactive oxygen species metabolism, (iii) small GTPases mediated signal transduction, (iv) circulatory and blood system processes, and (v) ER stress response (Table S1 and supplemental data 2). By its turn, cluster 2 comprises biological processes like: (i) hemopoiesis and leukocyte differentiation, (ii) positive regulation of JAK-STAT cascade, and (iii) ER-nucleus signaling pathway (Table S2 and supplementary data 2). As expected, proteins that could not be classified into any cluster were also identified in the network (Figure S2). Taking into account the data gathered from this initial systems biology analysis, we prompted to get more informations about the major nodes involved in the major PDI- 3
4 associated PPPI network using network centralities. Network centralities allow us to identify nodes (and the consequent biological processes) that have a relevant position in the overall network architecture (7) and many network centralities have been developed to evaluate the importance of a node for a given network, e.g., node degree, betweenness, and eigenvector measures (7). Centralities have been recently applied to quantify the centrality and prestige of actors in social networks (7) and to understand the structure and properties of complex biological, technological and infrastructural networks (6,7). Many of the nodes in a given network that show elevated values of centrality are important points of vulnerability, indicating that any attack to these nodes could introduce strong perturbations in the network. This graph principle has been exploited to identify proteins that are essential for an organism or that occupy a central position in a biological process, like bottleneck nodes (7-9) Bottleneck is a local topologic data that is defined as all nodes with high betweenness values and different nodes degrees, indicating that those nodes are central points that control the communication between other nodes within the network. Bottleneck also indicates all nodes that are between highly interconnected subgraph clusters, and removing a bottleneck could divide a network (10-12). Bottleneck nodes correspond to highly central proteins that connect several complexes or are peripheral members of central complexes, being important communication points between two complexes (10). Mostly of bottleneck nodes tend to be essential proteins in a network (10). The centrality analysis of major PDI-associated PPPI network (Figure S2) indicated the presence of 50 bottleneck nodes with different scores (Figure 4), corresponding to approximately 31% of all nodes present in the PPPI network. Interestingly, the bottlenecks with high score values correspond to proteins associated to small GTPases signaling processes, NADPH oxidases, intracellular signaling, and PDIA2 (Figure 4) 4
5 SUPPLEMENTAL REFERENCES 1. Shannon, P., Markiel, A., Ozier, O., Baliga, N. S., Wang, J. T., Ramage, D., Amin, N., Schwikowski, B., and Ideker, T. (2003) Genome research 13, Bader, G. D., and Hogue, C. W. (2003) BMC bioinformatics 4, 2 3. Lin, C. Y., Chin, C. H., Wu, H. H., Chen, S. H., Ho, C. W., and Ko, M. T. (2008) Nucleic acids research 36, W Maere, S., Heymans, K., and Kuiper, M. (2005) Bioinformatics 21, Rivals, I., Personnaz, L., Taing, L., and Potier, M. C. (2007) Bioinformatics 23, Benjamini, Y., and Hochberg, Y. (1995) J Roy Stat Soc B Met 57, Borgatti, S. P. (2005) Soc Networks 27, Estrada, E. (2006) Proteomics 6, Estrada, E., and Hatano, N. (2010) Physica A 389, Yu, H., Kim, P. M., Sprecher, E., Trifonov, V., and Gerstein, M. (2007) PLoS computational biology 3, e Newman, M. E. J. (2005) Soc Networks 27, Girvan, M., and Newman, M. E. J. (2002) Proceedings of the National Academy of Sciences of the United States of America 99,
6 SUPPLEMENTAL TABLES Table S1. Specific gene ontology (GO) classes derived from physical protein-protein interaction (PPPI) network observed in the cluster 1. Biologic Process GO classify p a value Corrected p value b k c f d Intracellular signal transduction Superoxide metabolic process Oxygen and reactive oxygen species metabolic process Small GTPase mediated signal transduction Ras protein signal transduction MAPKKK cascade Rho protein signal transduction Response to hydrogen peroxide Circulatory system process Blood circulation ER stress response Response to endoplasmic reticulum stress a p values calculated by the hypergeometric distribution of one ontology class visualized in the network. b Calculated values based on p values obtained after FDR was applied. c Total number of proteins found in the network which belong to a gene ontology. d Total number of proteins that belong to a specific gene ontology. 6
7 Table S2. Specific gene ontology (GO) classes derived from physical protein-protein interaction (PPPI) of unclustered proteins subnetwork. Biologic Process GO classify p a value Corrected p value b k c f d Hemopoiesis Leukocyte differentiation Hemopoietic or lymphoid organ development Positive regulation of JAK-STAT cascade ER-nucleus signaling pathway a p values calculated by the hypergeometric distribution of one ontology class visualized in the network. b Calculated values based on p values obtained after FDR was applied. c Total number of proteins found in the network which belong to a gene ontology. d Total number of proteins that belong to a specific gene ontology. 7
8 SUPPLEMENTAL FIGURE LEGENDS Figure S1. Effects of Diphenyleneiodonium on VSMC migration and ROS production. A) VSMC migration, analyzed by Boyden chamber assay, in VSMC exposed or not to PDGF (100ng/ml); B) Whole-cell production of superoxide (followed by the 2-hydroxyethidium signal, EOH) and other oxidants (followed by the ethidium signal, E) at baseline or after PDGF (100ng/ml; 2h). Figure S2. A physical protein-protein interaction (PPPI) network obtained from human PDIA2 interactomic data. The subnetworks and unclustered nodes that compose PPPI network are indicated by nodes with different shapes (inset). Bottleneck nodes are represent in the PPPI network by a color scale that indicates its bottleneck score value (from highest to the lowest value; see figure inset and supplementary material for additional informations). Figure S3. Subnetwork of proteins (cluster 1) associated to intracellular signal transduction, reactive oxygen species metabolism, small GTPases mediated signal transduction, circulatory and blood system processes, and ER stress response. Bottleneck nodes are represent in the cluster 1 by a color scale that indicates its bottleneck score value (from highest to the lowest value; see figure inset and supplementary material 2 for additional informations). Figure S4. Subnetwork of proteins (cluster 2) associated to hemopoiesis and leukocyte differentiation, positive regulation of JAK-STAT cascade, and ER-nucleus signaling pathway. Bottleneck nodes are represent in the cluster 2 by a color scale that indicates its bottleneck score value (from highest to the lowest value; see figure inset and supplementary material 2 for additional informations). 8
9 SUPPLEMENTAL FIGURES Supplemental Figure S1 9
10 Supplemental Figure S2. 10
11 Supplemental Figure S3 11
12 Supplemental Figure S4. 12
PNmerger: a Cytoscape plugin to merge biological pathways and protein interaction networks
 PNmerger: a Cytoscape plugin to merge biological pathways and protein interaction networks http://www.hupo.org.cn/pnmerger Fuchu He E-mail: hefc@nic.bmi.ac.cn Tel: 86-10-68171208 FAX: 86-10-68214653 Yunping
PNmerger: a Cytoscape plugin to merge biological pathways and protein interaction networks http://www.hupo.org.cn/pnmerger Fuchu He E-mail: hefc@nic.bmi.ac.cn Tel: 86-10-68171208 FAX: 86-10-68214653 Yunping
Networks & pathways. Hedi Peterson MTAT Bioinformatics
 Networks & pathways Hedi Peterson (peterson@quretec.com) MTAT.03.239 Bioinformatics 03.11.2010 Networks are graphs Nodes Edges Edges Directed, undirected, weighted Nodes Genes Proteins Metabolites Enzymes
Networks & pathways Hedi Peterson (peterson@quretec.com) MTAT.03.239 Bioinformatics 03.11.2010 Networks are graphs Nodes Edges Edges Directed, undirected, weighted Nodes Genes Proteins Metabolites Enzymes
Evidence for dynamically organized modularity in the yeast protein-protein interaction network
 Evidence for dynamically organized modularity in the yeast protein-protein interaction network Sari Bombino Helsinki 27.3.2007 UNIVERSITY OF HELSINKI Department of Computer Science Seminar on Computational
Evidence for dynamically organized modularity in the yeast protein-protein interaction network Sari Bombino Helsinki 27.3.2007 UNIVERSITY OF HELSINKI Department of Computer Science Seminar on Computational
Structure and Centrality of the Largest Fully Connected Cluster in Protein-Protein Interaction Networks
 22 International Conference on Environment Science and Engieering IPCEE vol.3 2(22) (22)ICSIT Press, Singapoore Structure and Centrality of the Largest Fully Connected Cluster in Protein-Protein Interaction
22 International Conference on Environment Science and Engieering IPCEE vol.3 2(22) (22)ICSIT Press, Singapoore Structure and Centrality of the Largest Fully Connected Cluster in Protein-Protein Interaction
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Networks. Can (John) Bruce Keck Founda7on Biotechnology Lab Bioinforma7cs Resource
 Networks Can (John) Bruce Keck Founda7on Biotechnology Lab Bioinforma7cs Resource Networks in biology Protein-Protein Interaction Network of Yeast Transcriptional regulatory network of E.coli Experimental
Networks Can (John) Bruce Keck Founda7on Biotechnology Lab Bioinforma7cs Resource Networks in biology Protein-Protein Interaction Network of Yeast Transcriptional regulatory network of E.coli Experimental
Comparative Network Analysis
 Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Comparison of Human Protein-Protein Interaction Maps
 Comparison of Human Protein-Protein Interaction Maps Matthias E. Futschik 1, Gautam Chaurasia 1,2, Erich Wanker 2 and Hanspeter Herzel 1 1 Institute for Theoretical Biology, Charité, Humboldt-Universität
Comparison of Human Protein-Protein Interaction Maps Matthias E. Futschik 1, Gautam Chaurasia 1,2, Erich Wanker 2 and Hanspeter Herzel 1 1 Institute for Theoretical Biology, Charité, Humboldt-Universität
Francisco M. Couto Mário J. Silva Pedro Coutinho
 Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
Analysis and visualization of protein-protein interactions. Olga Vitek Assistant Professor Statistics and Computer Science
 1 Analysis and visualization of protein-protein interactions Olga Vitek Assistant Professor Statistics and Computer Science 2 Outline 1. Protein-protein interactions 2. Using graph structures to study
1 Analysis and visualization of protein-protein interactions Olga Vitek Assistant Professor Statistics and Computer Science 2 Outline 1. Protein-protein interactions 2. Using graph structures to study
Lecture 3: A basic statistical concept
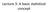 Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
A Multiobjective GO based Approach to Protein Complex Detection
 Available online at www.sciencedirect.com Procedia Technology 4 (2012 ) 555 560 C3IT-2012 A Multiobjective GO based Approach to Protein Complex Detection Sumanta Ray a, Moumita De b, Anirban Mukhopadhyay
Available online at www.sciencedirect.com Procedia Technology 4 (2012 ) 555 560 C3IT-2012 A Multiobjective GO based Approach to Protein Complex Detection Sumanta Ray a, Moumita De b, Anirban Mukhopadhyay
Proteomics Systems Biology
 Dr. Sanjeeva Srivastava IIT Bombay Proteomics Systems Biology IIT Bombay 2 1 DNA Genomics RNA Transcriptomics Global Cellular Protein Proteomics Global Cellular Metabolite Metabolomics Global Cellular
Dr. Sanjeeva Srivastava IIT Bombay Proteomics Systems Biology IIT Bombay 2 1 DNA Genomics RNA Transcriptomics Global Cellular Protein Proteomics Global Cellular Metabolite Metabolomics Global Cellular
Robust Community Detection Methods with Resolution Parameter for Complex Detection in Protein Protein Interaction Networks
 Robust Community Detection Methods with Resolution Parameter for Complex Detection in Protein Protein Interaction Networks Twan van Laarhoven and Elena Marchiori Institute for Computing and Information
Robust Community Detection Methods with Resolution Parameter for Complex Detection in Protein Protein Interaction Networks Twan van Laarhoven and Elena Marchiori Institute for Computing and Information
Supplementary methods
 Supplementary methods The rxncon language The network reconstruction was performed using the reaction-contingency (rxncon) language (1). The language is based on the separation of two distinct classes
Supplementary methods The rxncon language The network reconstruction was performed using the reaction-contingency (rxncon) language (1). The language is based on the separation of two distinct classes
Supplemental table S7.
 Supplemental table S7. GO terms significantly enriched in significantly up-regulated genes of the microarray. K: number of genes from the input cluster in the given category. F: number of total genes in
Supplemental table S7. GO terms significantly enriched in significantly up-regulated genes of the microarray. K: number of genes from the input cluster in the given category. F: number of total genes in
86 Part 4 SUMMARY INTRODUCTION
 86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
86 Part 4 Chapter # AN INTEGRATION OF THE DESCRIPTIONS OF GENE NETWORKS AND THEIR MODELS PRESENTED IN SIGMOID (CELLERATOR) AND GENENET Podkolodny N.L. *1, 2, Podkolodnaya N.N. 1, Miginsky D.S. 1, Poplavsky
Bioinformatics I. CPBS 7711 October 29, 2015 Protein interaction networks. Debra Goldberg
 Bioinformatics I CPBS 7711 October 29, 2015 Protein interaction networks Debra Goldberg debra@colorado.edu Overview Networks, protein interaction networks (PINs) Network models What can we learn from PINs
Bioinformatics I CPBS 7711 October 29, 2015 Protein interaction networks Debra Goldberg debra@colorado.edu Overview Networks, protein interaction networks (PINs) Network models What can we learn from PINs
Overview. Overview. Social networks. What is a network? 10/29/14. Bioinformatics I. Networks are everywhere! Introduction to Networks
 Bioinformatics I Overview CPBS 7711 October 29, 2014 Protein interaction networks Debra Goldberg debra@colorado.edu Networks, protein interaction networks (PINs) Network models What can we learn from PINs
Bioinformatics I Overview CPBS 7711 October 29, 2014 Protein interaction networks Debra Goldberg debra@colorado.edu Networks, protein interaction networks (PINs) Network models What can we learn from PINs
BMD645. Integration of Omics
 BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
BMD645 Integration of Omics Shu-Jen Chen, Chang Gung University Dec. 11, 2009 1 Traditional Biology vs. Systems Biology Traditional biology : Single genes or proteins Systems biology: Simultaneously study
Written Exam 15 December Course name: Introduction to Systems Biology Course no
 Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs
 Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs Xiao Chang 1,#, Tao Xu 2,#, Yun Li 3, Kai Wang 1,4,5,* 1 Zilkha Neurogenetic Institute,
Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs Xiao Chang 1,#, Tao Xu 2,#, Yun Li 3, Kai Wang 1,4,5,* 1 Zilkha Neurogenetic Institute,
Evolutionary Analysis of Functional Modules in Dynamic PPI Networks
 Evolutionary Analysis of Functional Modules in Dynamic PPI Networks ABSTRACT Nan Du Computer Science and Engineering Department nandu@buffalo.edu Jing Gao Computer Science and Engineering Department jing@buffalo.edu
Evolutionary Analysis of Functional Modules in Dynamic PPI Networks ABSTRACT Nan Du Computer Science and Engineering Department nandu@buffalo.edu Jing Gao Computer Science and Engineering Department jing@buffalo.edu
Protein function prediction via analysis of interactomes
 Protein function prediction via analysis of interactomes Elena Nabieva Mona Singh Department of Computer Science & Lewis-Sigler Institute for Integrative Genomics January 22, 2008 1 Introduction Genome
Protein function prediction via analysis of interactomes Elena Nabieva Mona Singh Department of Computer Science & Lewis-Sigler Institute for Integrative Genomics January 22, 2008 1 Introduction Genome
Graph Alignment and Biological Networks
 Graph Alignment and Biological Networks Johannes Berg http://www.uni-koeln.de/ berg Institute for Theoretical Physics University of Cologne Germany p.1/12 Networks in molecular biology New large-scale
Graph Alignment and Biological Networks Johannes Berg http://www.uni-koeln.de/ berg Institute for Theoretical Physics University of Cologne Germany p.1/12 Networks in molecular biology New large-scale
1 GO: regulation of cell size E-04 2 GO: negative regulation of cell growth GO:
 Table S2: The biological modulated by mir-5701 Sr. No Term Id 1 Term Name 2 Hit Gene Number 3 P-Value 4 1 GO:0008361 regulation of cell size 9 4.37E-04 2 GO:0030308 negative regulation of cell growth 8
Table S2: The biological modulated by mir-5701 Sr. No Term Id 1 Term Name 2 Hit Gene Number 3 P-Value 4 1 GO:0008361 regulation of cell size 9 4.37E-04 2 GO:0030308 negative regulation of cell growth 8
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 SUMOylation of proteins changes drastically upon heat shock, MG-132 treatment and PR-619 treatment. (a) Schematic overview of all SUMOylation proteins identified to be differentially
Supplementary Figure 1 SUMOylation of proteins changes drastically upon heat shock, MG-132 treatment and PR-619 treatment. (a) Schematic overview of all SUMOylation proteins identified to be differentially
Cellular Systems Biology or Biological Network Analysis
 Cellular Systems Biology or Biological Network Analysis Joel S. Bader Department of Biomedical Engineering Johns Hopkins University (c) 2012 December 4, 2012 1 Preface Cells are systems. Standard engineering
Cellular Systems Biology or Biological Network Analysis Joel S. Bader Department of Biomedical Engineering Johns Hopkins University (c) 2012 December 4, 2012 1 Preface Cells are systems. Standard engineering
Supplementary text for the section Interactions conserved across species: can one select the conserved interactions?
 1 Supporting Information: What Evidence is There for the Homology of Protein-Protein Interactions? Anna C. F. Lewis, Nick S. Jones, Mason A. Porter, Charlotte M. Deane Supplementary text for the section
1 Supporting Information: What Evidence is There for the Homology of Protein-Protein Interactions? Anna C. F. Lewis, Nick S. Jones, Mason A. Porter, Charlotte M. Deane Supplementary text for the section
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/8/e1500527/dc1 Supplementary Materials for A phylogenomic data-driven exploration of viral origins and evolution The PDF file includes: Arshan Nasir and Gustavo
advances.sciencemag.org/cgi/content/full/1/8/e1500527/dc1 Supplementary Materials for A phylogenomic data-driven exploration of viral origins and evolution The PDF file includes: Arshan Nasir and Gustavo
Proteomics. Yeast two hybrid. Proteomics - PAGE techniques. Data obtained. What is it?
 Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Identification of protein complexes from multi-relationship protein interaction networks
 Li et al. Human Genomics 2016, 10(Suppl 2):17 DOI 10.1186/s40246-016-0069-z RESEARCH Identification of protein complexes from multi-relationship protein interaction networks Xueyong Li 1,2, Jianxin Wang
Li et al. Human Genomics 2016, 10(Suppl 2):17 DOI 10.1186/s40246-016-0069-z RESEARCH Identification of protein complexes from multi-relationship protein interaction networks Xueyong Li 1,2, Jianxin Wang
Protein Complex Identification by Supervised Graph Clustering
 Protein Complex Identification by Supervised Graph Clustering Yanjun Qi 1, Fernanda Balem 2, Christos Faloutsos 1, Judith Klein- Seetharaman 1,2, Ziv Bar-Joseph 1 1 School of Computer Science, Carnegie
Protein Complex Identification by Supervised Graph Clustering Yanjun Qi 1, Fernanda Balem 2, Christos Faloutsos 1, Judith Klein- Seetharaman 1,2, Ziv Bar-Joseph 1 1 School of Computer Science, Carnegie
Computational Network Biology Biostatistics & Medical Informatics 826 Fall 2018
 Computational Network Biology Biostatistics & Medical Informatics 826 Fall 2018 Sushmita Roy sroy@biostat.wisc.edu https://compnetbiocourse.discovery.wisc.edu Sep 6 th 2018 Goals for today Administrivia
Computational Network Biology Biostatistics & Medical Informatics 826 Fall 2018 Sushmita Roy sroy@biostat.wisc.edu https://compnetbiocourse.discovery.wisc.edu Sep 6 th 2018 Goals for today Administrivia
GRAPH-THEORETICAL COMPARISON REVEALS STRUCTURAL DIVERGENCE OF HUMAN PROTEIN INTERACTION NETWORKS
 141 GRAPH-THEORETICAL COMPARISON REVEALS STRUCTURAL DIVERGENCE OF HUMAN PROTEIN INTERACTION NETWORKS MATTHIAS E. FUTSCHIK 1 ANNA TSCHAUT 2 m.futschik@staff.hu-berlin.de tschaut@zedat.fu-berlin.de GAUTAM
141 GRAPH-THEORETICAL COMPARISON REVEALS STRUCTURAL DIVERGENCE OF HUMAN PROTEIN INTERACTION NETWORKS MATTHIAS E. FUTSCHIK 1 ANNA TSCHAUT 2 m.futschik@staff.hu-berlin.de tschaut@zedat.fu-berlin.de GAUTAM
Gene Ontology. Shifra Ben-Dor. Weizmann Institute of Science
 Gene Ontology Shifra Ben-Dor Weizmann Institute of Science Outline of Session What is GO (Gene Ontology)? What tools do we use to work with it? Combination of GO with other analyses What is Ontology? 1700s
Gene Ontology Shifra Ben-Dor Weizmann Institute of Science Outline of Session What is GO (Gene Ontology)? What tools do we use to work with it? Combination of GO with other analyses What is Ontology? 1700s
Bioinformatics 2. Yeast two hybrid. Proteomics. Proteomics
 GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
Towards Detecting Protein Complexes from Protein Interaction Data
 Towards Detecting Protein Complexes from Protein Interaction Data Pengjun Pei 1 and Aidong Zhang 1 Department of Computer Science and Engineering State University of New York at Buffalo Buffalo NY 14260,
Towards Detecting Protein Complexes from Protein Interaction Data Pengjun Pei 1 and Aidong Zhang 1 Department of Computer Science and Engineering State University of New York at Buffalo Buffalo NY 14260,
MTopGO: a tool for module identification in PPI Networks
 MTopGO: a tool for module identification in PPI Networks Danila Vella 1,2, Simone Marini 3,4, Francesca Vitali 5,6,7, Riccardo Bellazzi 1,4 1 Clinical Scientific Institute Maugeri, Pavia, Italy, 2 Department
MTopGO: a tool for module identification in PPI Networks Danila Vella 1,2, Simone Marini 3,4, Francesca Vitali 5,6,7, Riccardo Bellazzi 1,4 1 Clinical Scientific Institute Maugeri, Pavia, Italy, 2 Department
Context dependent visualization of protein function
 Article III Context dependent visualization of protein function In: Juho Rousu, Samuel Kaski and Esko Ukkonen (eds.). Probabilistic Modeling and Machine Learning in Structural and Systems Biology. 2006,
Article III Context dependent visualization of protein function In: Juho Rousu, Samuel Kaski and Esko Ukkonen (eds.). Probabilistic Modeling and Machine Learning in Structural and Systems Biology. 2006,
Predicting Protein Functions and Domain Interactions from Protein Interactions
 Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Lecture 10: May 19, High-Throughput technologies for measuring proteinprotein
 Analysis of Gene Expression Data Spring Semester, 2005 Lecture 10: May 19, 2005 Lecturer: Roded Sharan Scribe: Daniela Raijman and Igor Ulitsky 10.1 Protein Interaction Networks In the past we have discussed
Analysis of Gene Expression Data Spring Semester, 2005 Lecture 10: May 19, 2005 Lecturer: Roded Sharan Scribe: Daniela Raijman and Igor Ulitsky 10.1 Protein Interaction Networks In the past we have discussed
Research Article Prediction of Protein-Protein Interactions Related to Protein Complexes Based on Protein Interaction Networks
 BioMed Research International Volume 2015, Article ID 259157, 9 pages http://dx.doi.org/10.1155/2015/259157 Research Article Prediction of Protein-Protein Interactions Related to Protein Complexes Based
BioMed Research International Volume 2015, Article ID 259157, 9 pages http://dx.doi.org/10.1155/2015/259157 Research Article Prediction of Protein-Protein Interactions Related to Protein Complexes Based
Systems biology and biological networks
 Systems Biology Workshop Systems biology and biological networks Center for Biological Sequence Analysis Networks in electronics Radio kindly provided by Lazebnik, Cancer Cell, 2002 Systems Biology Workshop,
Systems Biology Workshop Systems biology and biological networks Center for Biological Sequence Analysis Networks in electronics Radio kindly provided by Lazebnik, Cancer Cell, 2002 Systems Biology Workshop,
Interaction Network Analysis
 CSI/BIF 5330 Interaction etwork Analsis Young-Rae Cho Associate Professor Department of Computer Science Balor Universit Biological etworks Definition Maps of biochemical reactions, interactions, regulations
CSI/BIF 5330 Interaction etwork Analsis Young-Rae Cho Associate Professor Department of Computer Science Balor Universit Biological etworks Definition Maps of biochemical reactions, interactions, regulations
2 GENE FUNCTIONAL SIMILARITY. 2.1 Semantic values of GO terms
 Bioinformatics Advance Access published March 7, 2007 The Author (2007). Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oxfordjournals.org
Bioinformatics Advance Access published March 7, 2007 The Author (2007). Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oxfordjournals.org
Basic modeling approaches for biological systems. Mahesh Bule
 Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Lecture Notes for Fall Network Modeling. Ernest Fraenkel
 Lecture Notes for 20.320 Fall 2012 Network Modeling Ernest Fraenkel In this lecture we will explore ways in which network models can help us to understand better biological data. We will explore how networks
Lecture Notes for 20.320 Fall 2012 Network Modeling Ernest Fraenkel In this lecture we will explore ways in which network models can help us to understand better biological data. We will explore how networks
Preface. Contributors
 CONTENTS Foreword Preface Contributors PART I INTRODUCTION 1 1 Networks in Biology 3 Björn H. Junker 1.1 Introduction 3 1.2 Biology 101 4 1.2.1 Biochemistry and Molecular Biology 4 1.2.2 Cell Biology 6
CONTENTS Foreword Preface Contributors PART I INTRODUCTION 1 1 Networks in Biology 3 Björn H. Junker 1.1 Introduction 3 1.2 Biology 101 4 1.2.1 Biochemistry and Molecular Biology 4 1.2.2 Cell Biology 6
Modular organization of protein interaction networks
 BIOINFORMATICS ORIGINAL PAPER Vol. 23 no. 2 2007, pages 207 214 doi:10.1093/bioinformatics/btl562 Systems biology Modular organization of protein interaction networks Feng Luo 1,3,, Yunfeng Yang 2, Chin-Fu
BIOINFORMATICS ORIGINAL PAPER Vol. 23 no. 2 2007, pages 207 214 doi:10.1093/bioinformatics/btl562 Systems biology Modular organization of protein interaction networks Feng Luo 1,3,, Yunfeng Yang 2, Chin-Fu
Systematic prediction of gene function in Arabidopsis thaliana using a probabilistic functional gene network
 Systematic prediction of gene function in Arabidopsis thaliana using a probabilistic functional gene network Sohyun Hwang 1, Seung Y Rhee 2, Edward M Marcotte 3,4 & Insuk Lee 1 protocol 1 Department of
Systematic prediction of gene function in Arabidopsis thaliana using a probabilistic functional gene network Sohyun Hwang 1, Seung Y Rhee 2, Edward M Marcotte 3,4 & Insuk Lee 1 protocol 1 Department of
hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference
 CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
Organization of Physical Interactomes as Uncovered by Network Schemas
 Organization of Physical Interactomes as Uncovered by Network Schemas Eric Banks., Elena Nabieva., Bernard Chazelle, Mona Singh* Department of Computer Science & Lewis-Sigler Institute for Integrative
Organization of Physical Interactomes as Uncovered by Network Schemas Eric Banks., Elena Nabieva., Bernard Chazelle, Mona Singh* Department of Computer Science & Lewis-Sigler Institute for Integrative
Fine-scale dissection of functional protein network. organization by dynamic neighborhood analysis
 Fine-scale dissection of functional protein network organization by dynamic neighborhood analysis Kakajan Komurov 1, Mehmet H. Gunes 2, Michael A. White 1 1 Department of Cell Biology, University of Texas
Fine-scale dissection of functional protein network organization by dynamic neighborhood analysis Kakajan Komurov 1, Mehmet H. Gunes 2, Michael A. White 1 1 Department of Cell Biology, University of Texas
Androgen-independent prostate cancer
 The following tutorial walks through the identification of biological themes in a microarray dataset examining androgen-independent. Visit the GeneSifter Data Center (www.genesifter.net/web/datacenter.html)
The following tutorial walks through the identification of biological themes in a microarray dataset examining androgen-independent. Visit the GeneSifter Data Center (www.genesifter.net/web/datacenter.html)
Supplemental material
 Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Supplemental material THE JOURNAL OF CELL BIOLOGY Mourier et al., http://www.jcb.org/cgi/content/full/jcb.201411100/dc1 Figure S1. Size and mitochondrial content in Mfn1 and Mfn2 knockout hearts. (A) Body
Assessing Significance of Connectivity and. Conservation in Protein Interaction Networks
 Assessing Significance of Connectivity and Conservation in Protein Interaction Networks Mehmet Koyutürk, Wojciech Szpankowski, and Ananth Grama Department of Computer Sciences, Purdue University, West
Assessing Significance of Connectivity and Conservation in Protein Interaction Networks Mehmet Koyutürk, Wojciech Szpankowski, and Ananth Grama Department of Computer Sciences, Purdue University, West
Network Biology: Understanding the cell s functional organization. Albert-László Barabási Zoltán N. Oltvai
 Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling
 Lethality and centrality in protein networks Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling molecules, or building blocks of cells and microorganisms.
Lethality and centrality in protein networks Cell biology traditionally identifies proteins based on their individual actions as catalysts, signaling molecules, or building blocks of cells and microorganisms.
Inferring Transcriptional Regulatory Networks from Gene Expression Data II
 Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
MTGO: PPI Network Analysis Via Topological and Functional Module Identification
 www.nature.com/scientificreports Received: 1 November 2017 Accepted: 28 February 2018 Published: xx xx xxxx OPEN MTGO: PPI Network Analysis Via Topological and Functional Module Identification Danila Vella
www.nature.com/scientificreports Received: 1 November 2017 Accepted: 28 February 2018 Published: xx xx xxxx OPEN MTGO: PPI Network Analysis Via Topological and Functional Module Identification Danila Vella
Gene Ontology and overrepresentation analysis
 Gene Ontology and overrepresentation analysis Kjell Petersen J Express Microarray analysis course Oslo December 2009 Presentation adapted from Endre Anderssen and Vidar Beisvåg NMC Trondheim Overview How
Gene Ontology and overrepresentation analysis Kjell Petersen J Express Microarray analysis course Oslo December 2009 Presentation adapted from Endre Anderssen and Vidar Beisvåg NMC Trondheim Overview How
Profiling Human Cell/Tissue Specific Gene Regulation Networks
 Profiling Human Cell/Tissue Specific Gene Regulation Networks Louxin Zhang Department of Mathematics National University of Singapore matzlx@nus.edu.sg Network Biology < y 1 t, y 2 t,, y k t > u 1 u 2
Profiling Human Cell/Tissue Specific Gene Regulation Networks Louxin Zhang Department of Mathematics National University of Singapore matzlx@nus.edu.sg Network Biology < y 1 t, y 2 t,, y k t > u 1 u 2
Comparing transcription factor regulatory networks of human cell types. The Protein Network Workshop June 8 12, 2015
 Comparing transcription factor regulatory networks of human cell types The Protein Network Workshop June 8 12, 2015 KWOK-PUI CHOI Dept of Statistics & Applied Probability, Dept of Mathematics, NUS OUTLINE
Comparing transcription factor regulatory networks of human cell types The Protein Network Workshop June 8 12, 2015 KWOK-PUI CHOI Dept of Statistics & Applied Probability, Dept of Mathematics, NUS OUTLINE
Correlation Networks
 QuickTime decompressor and a are needed to see this picture. Correlation Networks Analysis of Biological Networks April 24, 2010 Correlation Networks - Analysis of Biological Networks 1 Review We have
QuickTime decompressor and a are needed to see this picture. Correlation Networks Analysis of Biological Networks April 24, 2010 Correlation Networks - Analysis of Biological Networks 1 Review We have
Comparison of Protein-Protein Interaction Confidence Assignment Schemes
 Comparison of Protein-Protein Interaction Confidence Assignment Schemes Silpa Suthram 1, Tomer Shlomi 2, Eytan Ruppin 2, Roded Sharan 2, and Trey Ideker 1 1 Department of Bioengineering, University of
Comparison of Protein-Protein Interaction Confidence Assignment Schemes Silpa Suthram 1, Tomer Shlomi 2, Eytan Ruppin 2, Roded Sharan 2, and Trey Ideker 1 1 Department of Bioengineering, University of
Why Do Hubs in the Yeast Protein Interaction Network Tend To Be Essential: Reexamining the Connection between the Network Topology and Essentiality
 Why Do Hubs in the Yeast Protein Interaction Network Tend To Be Essential: Reexamining the Connection between the Network Topology and Essentiality Elena Zotenko 1, Julian Mestre 1, Dianne P. O Leary 2,3,
Why Do Hubs in the Yeast Protein Interaction Network Tend To Be Essential: Reexamining the Connection between the Network Topology and Essentiality Elena Zotenko 1, Julian Mestre 1, Dianne P. O Leary 2,3,
FROM UNCERTAIN PROTEIN INTERACTION NETWORKS TO SIGNALING PATHWAYS THROUGH INTENSIVE COLOR CODING
 FROM UNCERTAIN PROTEIN INTERACTION NETWORKS TO SIGNALING PATHWAYS THROUGH INTENSIVE COLOR CODING Haitham Gabr, Alin Dobra and Tamer Kahveci CISE Department, University of Florida, Gainesville, FL 32611,
FROM UNCERTAIN PROTEIN INTERACTION NETWORKS TO SIGNALING PATHWAYS THROUGH INTENSIVE COLOR CODING Haitham Gabr, Alin Dobra and Tamer Kahveci CISE Department, University of Florida, Gainesville, FL 32611,
Hands-On Nine The PAX6 Gene and Protein
 Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
The Role of Network Science in Biology and Medicine. Tiffany J. Callahan Computational Bioscience Program Hunter/Kahn Labs
 The Role of Network Science in Biology and Medicine Tiffany J. Callahan Computational Bioscience Program Hunter/Kahn Labs Network Analysis Working Group 09.28.2017 Network-Enabled Wisdom (NEW) empirically
The Role of Network Science in Biology and Medicine Tiffany J. Callahan Computational Bioscience Program Hunter/Kahn Labs Network Analysis Working Group 09.28.2017 Network-Enabled Wisdom (NEW) empirically
Analysis of Biological Networks: Network Robustness and Evolution
 Analysis of Biological Networks: Network Robustness and Evolution Lecturer: Roded Sharan Scribers: Sasha Medvedovsky and Eitan Hirsh Lecture 14, February 2, 2006 1 Introduction The chapter is divided into
Analysis of Biological Networks: Network Robustness and Evolution Lecturer: Roded Sharan Scribers: Sasha Medvedovsky and Eitan Hirsh Lecture 14, February 2, 2006 1 Introduction The chapter is divided into
Detecting temporal protein complexes from dynamic protein-protein interaction networks
 Detecting temporal protein complexes from dynamic protein-protein interaction networks Le Ou-Yang, Dao-Qing Dai, Xiao-Li Li, Min Wu, Xiao-Fei Zhang and Peng Yang 1 Supplementary Table Table S1: Comparative
Detecting temporal protein complexes from dynamic protein-protein interaction networks Le Ou-Yang, Dao-Qing Dai, Xiao-Li Li, Min Wu, Xiao-Fei Zhang and Peng Yang 1 Supplementary Table Table S1: Comparative
CSCI1950 Z Computa3onal Methods for Biology Lecture 24. Ben Raphael April 29, hgp://cs.brown.edu/courses/csci1950 z/ Network Mo3fs
 CSCI1950 Z Computa3onal Methods for Biology Lecture 24 Ben Raphael April 29, 2009 hgp://cs.brown.edu/courses/csci1950 z/ Network Mo3fs Subnetworks with more occurrences than expected by chance. How to
CSCI1950 Z Computa3onal Methods for Biology Lecture 24 Ben Raphael April 29, 2009 hgp://cs.brown.edu/courses/csci1950 z/ Network Mo3fs Subnetworks with more occurrences than expected by chance. How to
Supplementary Information 16
 Supplementary Information 16 Cellular Component % of Genes 50 45 40 35 30 25 20 15 10 5 0 human mouse extracellular other membranes plasma membrane cytosol cytoskeleton mitochondrion ER/Golgi translational
Supplementary Information 16 Cellular Component % of Genes 50 45 40 35 30 25 20 15 10 5 0 human mouse extracellular other membranes plasma membrane cytosol cytoskeleton mitochondrion ER/Golgi translational
NETWORK CLUSTERING METHODS
 1 NETWORK CLUSTERING METHODS Dr. Alioune Ngom School of Computer Science University of Windsor angom@uwindsor.ca Winter 2013 Why clustering? 2 A cluster is a group of related objects In biological nets,
1 NETWORK CLUSTERING METHODS Dr. Alioune Ngom School of Computer Science University of Windsor angom@uwindsor.ca Winter 2013 Why clustering? 2 A cluster is a group of related objects In biological nets,
Supplementary Figure 3
 Supplementary Figure 3 a 1 (i) (ii) (iii) (iv) (v) log P gene Q group, % ~ ε nominal 2 1 1 8 6 5 A B C D D' G J L M P R U + + ε~ A C B D D G JL M P R U -1 1 ε~ (vi) Z group 2 1 1 (vii) (viii) Z module
Supplementary Figure 3 a 1 (i) (ii) (iii) (iv) (v) log P gene Q group, % ~ ε nominal 2 1 1 8 6 5 A B C D D' G J L M P R U + + ε~ A C B D D G JL M P R U -1 1 ε~ (vi) Z group 2 1 1 (vii) (viii) Z module
CS612 - Algorithms in Bioinformatics
 Fall 2017 Databases and Protein Structure Representation October 2, 2017 Molecular Biology as Information Science > 12, 000 genomes sequenced, mostly bacterial (2013) > 5x10 6 unique sequences available
Fall 2017 Databases and Protein Structure Representation October 2, 2017 Molecular Biology as Information Science > 12, 000 genomes sequenced, mostly bacterial (2013) > 5x10 6 unique sequences available
Genome-wide multilevel spatial interactome model of rice
 Sino-German Workshop on Multiscale Spatial Computational Systems Biology, Beijing, Oct 8-12, 2015 Genome-wide multilevel spatial interactome model of rice Ming CHEN ( 陈铭 ) mchen@zju.edu.cn College of Life
Sino-German Workshop on Multiscale Spatial Computational Systems Biology, Beijing, Oct 8-12, 2015 Genome-wide multilevel spatial interactome model of rice Ming CHEN ( 陈铭 ) mchen@zju.edu.cn College of Life
An Efficient Algorithm for Protein-Protein Interaction Network Analysis to Discover Overlapping Functional Modules
 An Efficient Algorithm for Protein-Protein Interaction Network Analysis to Discover Overlapping Functional Modules Ying Liu 1 Department of Computer Science, Mathematics and Science, College of Professional
An Efficient Algorithm for Protein-Protein Interaction Network Analysis to Discover Overlapping Functional Modules Ying Liu 1 Department of Computer Science, Mathematics and Science, College of Professional
Integration of functional genomics data
 Integration of functional genomics data Laboratoire Bordelais de Recherche en Informatique (UMR) Centre de Bioinformatique de Bordeaux (Plateforme) Rennes Oct. 2006 1 Observations and motivations Genomics
Integration of functional genomics data Laboratoire Bordelais de Recherche en Informatique (UMR) Centre de Bioinformatique de Bordeaux (Plateforme) Rennes Oct. 2006 1 Observations and motivations Genomics
Protein-protein interaction networks Prof. Peter Csermely
 Protein-Protein Interaction Networks 1 Department of Medical Chemistry Semmelweis University, Budapest, Hungary www.linkgroup.hu csermely@eok.sote.hu Advantages of multi-disciplinarity Networks have general
Protein-Protein Interaction Networks 1 Department of Medical Chemistry Semmelweis University, Budapest, Hungary www.linkgroup.hu csermely@eok.sote.hu Advantages of multi-disciplinarity Networks have general
Deciphering regulatory networks by promoter sequence analysis
 Bioinformatics Workshop 2009 Interpreting Gene Lists from -omics Studies Deciphering regulatory networks by promoter sequence analysis Elodie Portales-Casamar University of British Columbia www.cisreg.ca
Bioinformatics Workshop 2009 Interpreting Gene Lists from -omics Studies Deciphering regulatory networks by promoter sequence analysis Elodie Portales-Casamar University of British Columbia www.cisreg.ca
Computational Structural Bioinformatics
 Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
Computational Structural Bioinformatics ECS129 Instructor: Patrice Koehl http://koehllab.genomecenter.ucdavis.edu/teaching/ecs129 koehl@cs.ucdavis.edu Learning curve Math / CS Biology/ Chemistry Pre-requisite
arxiv: v1 [q-bio.mn] 5 Feb 2008
![arxiv: v1 [q-bio.mn] 5 Feb 2008 arxiv: v1 [q-bio.mn] 5 Feb 2008](/thumbs/82/86837131.jpg) Uncovering Biological Network Function via Graphlet Degree Signatures Tijana Milenković and Nataša Pržulj Department of Computer Science, University of California, Irvine, CA 92697-3435, USA Technical
Uncovering Biological Network Function via Graphlet Degree Signatures Tijana Milenković and Nataša Pržulj Department of Computer Science, University of California, Irvine, CA 92697-3435, USA Technical
An introduction to SYSTEMS BIOLOGY
 An introduction to SYSTEMS BIOLOGY Paolo Tieri CNR Consiglio Nazionale delle Ricerche, Rome, Italy 10 February 2015 Universidade Federal de Minas Gerais, Belo Horizonte, Brasil Course outline Day 1: intro
An introduction to SYSTEMS BIOLOGY Paolo Tieri CNR Consiglio Nazionale delle Ricerche, Rome, Italy 10 February 2015 Universidade Federal de Minas Gerais, Belo Horizonte, Brasil Course outline Day 1: intro
Pathway Association Analysis Trey Ideker UCSD
 Pathway Association Analysis Trey Ideker UCSD A working network map of the cell Network evolutionary comparison / cross-species alignment to identify conserved modules The Working Map Network-based classification
Pathway Association Analysis Trey Ideker UCSD A working network map of the cell Network evolutionary comparison / cross-species alignment to identify conserved modules The Working Map Network-based classification
International Journal of Scientific & Engineering Research, Volume 6, Issue 2, February ISSN
 International Journal of Scientific & Engineering Research, Volume 6, Issue 2, February-2015 273 Analogizing And Investigating Some Applications of Metabolic Pathway Analysis Methods Gourav Mukherjee 1,
International Journal of Scientific & Engineering Research, Volume 6, Issue 2, February-2015 273 Analogizing And Investigating Some Applications of Metabolic Pathway Analysis Methods Gourav Mukherjee 1,
An ontology-based search engine for protein-protein interactions Byungkyu Park and Kyungsook Han*
 BMC Bioinformatics BioMed Central Research An ontology-based search engine for protein-protein interactions Byungkyu Park and Kyungsook Han* Open Access Address: School of Computer Science and Engineering,
BMC Bioinformatics BioMed Central Research An ontology-based search engine for protein-protein interactions Byungkyu Park and Kyungsook Han* Open Access Address: School of Computer Science and Engineering,
Discovering molecular pathways from protein interaction and ge
 Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
KEGGgraph: Application Examples
 KEGGgraph: Application Examples Jitao David Zhang April 30, 2018 Abstract In this vignette, we demonstrate the application of KEGGgraph as flexible module in analysis pipelines targeting heterogenous biological
KEGGgraph: Application Examples Jitao David Zhang April 30, 2018 Abstract In this vignette, we demonstrate the application of KEGGgraph as flexible module in analysis pipelines targeting heterogenous biological
Dynamic modeling and analysis of cancer cellular network motifs
 SUPPLEMENTARY MATERIAL 1: Dynamic modeling and analysis of cancer cellular network motifs Mathieu Cloutier 1 and Edwin Wang 1,2* 1. Computational Chemistry and Bioinformatics Group, Biotechnology Research
SUPPLEMENTARY MATERIAL 1: Dynamic modeling and analysis of cancer cellular network motifs Mathieu Cloutier 1 and Edwin Wang 1,2* 1. Computational Chemistry and Bioinformatics Group, Biotechnology Research
Supplementary Figure 1 The number of differentially expressed genes for uniparental males (green), uniparental females (yellow), biparental males
 Supplementary Figure 1 The number of differentially expressed genes for males (green), females (yellow), males (red), and females (blue) in caring vs. control comparisons in the caring gene set and the
Supplementary Figure 1 The number of differentially expressed genes for males (green), females (yellow), males (red), and females (blue) in caring vs. control comparisons in the caring gene set and the
Supplemental Material
 Supplemental Material Article title: Construction and comparison of gene co expression networks shows complex plant immune responses Author names and affiliation: Luis Guillermo Leal 1*, Email: lgleala@unal.edu.co
Supplemental Material Article title: Construction and comparison of gene co expression networks shows complex plant immune responses Author names and affiliation: Luis Guillermo Leal 1*, Email: lgleala@unal.edu.co
A taxonomy of visualization tasks for the analysis of biological pathway data
 The Author(s) BMC Bioinformatics 2016, 18(Suppl 2):21 DOI 10.1186/s12859-016-1443-5 RESEARCH A taxonomy of visualization tasks for the analysis of biological pathway data Paul Murray 1*,FintanMcGee 2 and
The Author(s) BMC Bioinformatics 2016, 18(Suppl 2):21 DOI 10.1186/s12859-016-1443-5 RESEARCH A taxonomy of visualization tasks for the analysis of biological pathway data Paul Murray 1*,FintanMcGee 2 and
Molecular Cell Biology 5068 In Class Exam 2 November 8, 2016
 Molecular Cell Biology 5068 In Class Exam 2 November 8, 2016 Exam Number: Please print your name: Instructions: Please write only on these pages, in the spaces allotted and not on the back. Write your
Molecular Cell Biology 5068 In Class Exam 2 November 8, 2016 Exam Number: Please print your name: Instructions: Please write only on these pages, in the spaces allotted and not on the back. Write your
EFFICIENT AND ROBUST PREDICTION ALGORITHMS FOR PROTEIN COMPLEXES USING GOMORY-HU TREES
 EFFICIENT AND ROBUST PREDICTION ALGORITHMS FOR PROTEIN COMPLEXES USING GOMORY-HU TREES A. MITROFANOVA*, M. FARACH-COLTON**, AND B. MISHRA* *New York University, Department of Computer Science, New York,
EFFICIENT AND ROBUST PREDICTION ALGORITHMS FOR PROTEIN COMPLEXES USING GOMORY-HU TREES A. MITROFANOVA*, M. FARACH-COLTON**, AND B. MISHRA* *New York University, Department of Computer Science, New York,
*Equal contribution Contact: (TT) 1 Department of Biomedical Engineering, the Engineering Faculty, Tel Aviv
 Supplementary of Complementary Post Transcriptional Regulatory Information is Detected by PUNCH-P and Ribosome Profiling Hadas Zur*,1, Ranen Aviner*,2, Tamir Tuller 1,3 1 Department of Biomedical Engineering,
Supplementary of Complementary Post Transcriptional Regulatory Information is Detected by PUNCH-P and Ribosome Profiling Hadas Zur*,1, Ranen Aviner*,2, Tamir Tuller 1,3 1 Department of Biomedical Engineering,
Research Article A Topological Description of Hubs in Amino Acid Interaction Networks
 Advances in Bioinformatics Volume 21, Article ID 257512, 9 pages doi:1.1155/21/257512 Research Article A Topological Description of Hubs in Amino Acid Interaction Networks Omar Gaci Le Havre University,
Advances in Bioinformatics Volume 21, Article ID 257512, 9 pages doi:1.1155/21/257512 Research Article A Topological Description of Hubs in Amino Acid Interaction Networks Omar Gaci Le Havre University,
Protein Interaction Mapping: Use of Osprey to map Survival of Motor Neuron Protein interactions
 Protein Interaction Mapping: Use of Osprey to map Survival of Motor Neuron Protein interactions Presented by: Meg Barnhart Computational Biosciences Arizona State University The Spinal Muscular Atrophy
Protein Interaction Mapping: Use of Osprey to map Survival of Motor Neuron Protein interactions Presented by: Meg Barnhart Computational Biosciences Arizona State University The Spinal Muscular Atrophy

 keâesme& Meer
keâesme& Meer