Supertree Algorithms for Ancestral Divergence Dates and Nested Taxa
|
|
|
- Trevor Lynch
- 5 years ago
- Views:
Transcription
1 Supertree Algorithms for Ancestral Divergence Dates and Nested Taxa Charles Semple 1, Philip Daniel 1, Wim Hordijk 1, Roderic D. M. Page 2, and Mike Steel 1 1 Biomathematics Research Centre, Department of Mathematics and Statistics, University of Canterbury, Christchurch, New Zealand 2 DEEB, IBLS, Graham Kerr Building, University of Glasgow, Glasgow G12 QP, United Kingdom Abstract. Motivation: Supertree methods have been often identified as a possible approach to the reconstruction of the Tree of Life. However, a limitation of such methods is that, typically, they use just leaf-labelled phylogenetic trees to infer the resulting supertree. Results: In this paper, we describe several new supertree algorithms that extend the allowable information that can be used for phylogenetic inference. These algorithms have been recently implemented and we describe here two illustrative applications. Availability: These new algorithms are freely available for application at Contact: c.semple@math.canterbury.ac.nz 1 Introduction Rooted phylogenetic trees are used in evolutionary biology to represent the ancestral history of a collection of present-day species. For example, ignoring the numerals, Fig. 1 shows a rooted phylogenetic tree where the labels a, b, c, d, e, and f represent the present-day species. A supertree is a rooted phylogenetic tree that is the result of combining a collection of smaller rooted phylogenetic trees on overlapping subsets of species. There are now many techniques ( supertree methods ) for constructing supertrees. However, for almost all of these methods, the input is restricted to leaf-labelled phylogenetic trees (without branch lengths or interior node labels or dates), in which case any other related information is ignored. In this paper, we describe several new, and recently implemented, supertree methods which extend the typical input to include some of this additional information. The new supertree methods divide into two groups depending upon the type of additional information being used as input. The first group allows the input to include ancestral divergence dates which may be either relative or explicit.
2 For example, in this group the input could include information such as whether one particular divergence event on one side of a tree occurred before or after a divergence event on the other side of the tree, or actual time estimates of certain divergence events. The second group of supertree algorithms takes as its input rooted trees in which some of the interior vertices as well as all of their leaves are labelled. This allows the inclusion of nested taxa in the input. All of the new supertree algorithms described in this paper have been implemented in Java (and are thus platform independent) and are available for applications at The implementation requires the input trees to be given in the Newick format 1. Any tree that is returned by these algorithms is also in the Newick format. All of the algorithms run quickly (that is, require just polynomial time) in the size of the input. The notation and terminology in this paper follows Semple and Steel (2003). The paper is organised as follows. Each of the new algorithms can be viewed as a variation of Build, one of the oldest supertree algorithms. Indeed, the general approach used in each of the algorithms is similar to that used by Build. This approach is outlined in the next section. In Sections 3 and 4, we describe the new algorithms for ancestral divergence dates and nested taxa, respectively, and include two applications of these algorithms to some data sets. We end this section by noting that the formal details of the algorithms described in this paper, including their correctness, are not included here as these details are to appear as chapters (Bryant et al., 2004; Daniel and Semple, 2004) of a forthcoming book on supertrees (Bininda-Emonds, 2004). 2 The Build Approach Originally designed for other purposes, Build (Aho et al., 191) is an exact algorithm in that it outputs a tree precisely if the input collection satisfies a particular compatibility criteria. A rooted phylogenetic tree T displays a rooted phylogenetic tree T if the label set X of T is a subset of the label set of T and, up to suppressing degree-two vertices, T is a refinement of the minimal rooted subtree of T that connects the labels in X. For the purposes of this paper, up to suppressing degree-two vertices essentially means overlooking degree-two vertices. A collection P of rooted phylogenetic trees is compatible 1 See for example
3 if there exists a rooted phylogenetic tree that displays each of the trees in P. Intuitively, P is compatible if there is a rooted phylogenetic tree that, up to polytomies, preserves all of the ancestral relationships described by the trees in P. In particular, if the most recent common ancestor of a and b is a descendant of the most recent common ancestor of a and c in a tree in P, then this relationship is also preserved in T. The algorithms we present in this paper follow the approach of Build. Each algorithm is exact and outputs a tree precisely if the input collection satisfies some particular compatibility criteria. Furthermore, like Build, the descriptions of the algorithms take the following form. Each algorithm attempts to construct a tree T that satisfies the compatibility criteria by constructing the set of clusters of T. In all cases, this is done by starting with the cluster that is the union of the labels of the trees in P and successively breaking it down into disjoint subclusters. How the clusters are broken down is determined by an associated graph at each iteration. For each algorithm, this graph as well as the process for breaking up clusters is different. The process continues provided the graph at each iteration satisfies some condition. Eventually, either (i) the algorithm outputs a tree that satisfies the compatibility criteria, or (ii) outputs a statement indicating that the input collection does not satisfy this criteria. 3 Supertree Algorithms for Ancestral Divergence Dates We first describe a supertree algorithm that incorporates relative divergence times. This algorithm is called RankedTree. An extension of this algorithm to include absolute divergence times or intervals on these times is also possible and this is mentioned at the end of this section. Essentially, RankedTree is an extension of Build and its input consists of rooted phylogenetic trees as well as information detailing the order in which the divergence events of certain different pairs of species occurred. We call the latter type of input a relative divergence date and such information is based, for example, on fossil data or molecular dating techniques. Formally, this type of input takes the form div(c, d) predates div(a, b) which is interpreted as, for species a, b, c, and d, the divergence of species c and d predates the divergence of species a and b. To include both types of input on a single supertree, we extend the concept of a rooted phylogenetic tree. A ranked phylogenetic tree T is a rooted phylogenetic tree in which the interior vertices are assigned a positive integer so that if v 1, v 2 are interior vertices and v 2 is a descendant of v 1, then the integer assigned to v 1 is less than the integer assigned to v 2. Such an assignment of positive integers is a
4 called a ranking of the interior vertices of T. An example of a ranked phylogenetic tree is shown in Fig. 1. Ranking the interior vertices of T in this way corresponds to an ordering of the speciation events associated to these vertices. Note that two different interior vertices may be assigned the same positive integer, in which case, it is inferred that there is no particular ordering on the associated speciation events a e d b c f Fig. 1. A ranked phylogenetic tree. A relative divergence date div(c, d) predates div(a, b) is preserved by a ranked phylogenetic tree T if a, b, c, d are leaf labels of T, and the rank assigned to the interior vertex of T corresponding to the most recent common ancestor of c and d is less than the rank assigned to the interior vertex of T corresponding to the most recent common ancestor of a and b. Thus, for example, the ranked phylogenetic tree shown in Fig. 1 preserves the relative divergence date div(e, b) predates div(c, f). A collection P of rooted phylogenetic trees and a collection D of relative divergence dates are compatible if there is a ranked phylogenetic tree T such that the discrete topology of T displays each of the trees in P and the ranking of the interior vertices of T preserves all of the relative divergence dates in D. The algorithm RankedTree decides whether or not collections of rooted phylogenetic trees and relative divergence dates are compatible. Furthermore, if these collections are compatible, then RankedTree returns a ranked phylogenetic tree that displays each of the rooted phylogenetic trees and preserves each of the relative divergence dates. To illustrate RankedTree, consider the two ranked phylogenetic trees shown in Fig. 2(a) (Janczewski et al., 1995) and (b) (Slattery et al., 1994), each of which is a phylogenetic tree of the cat family. The species labels are the 3-letter abbreviations used in these references. The branch lengths of the source trees have been translated into rankings and added to the interior vertices of these trees. (These branch lengths are also shown.) Observing that species LPA, PON, and CCR are common to both trees, the ranked phylogenetic tree shown in Fig. 2(c) is the result of applying RankedTree to these two trees. Note that the branch lengths of the resulting ranked phylogenetic tree do not reflect real time.
5 1 (a) PLE PPA PON PUN NNE PTI LCA LRU PMA PTE AJU PCO LSE CCA PAU FCA LPA 1 (b) LPA LWI OGU OGE LCO PON LTI CCR CCR LCA LRU PMA (c) 5 CCA PAU LSE AJU PCO PTE PLE PPA PON PUN NNE PTI 2 LTI FCA OGE LCO OGU LPA LWI CCR Fig. 2. An application of RankedTree. An extension of RankedTree allows for time bounds on speciation events as well as rooted phylogenetic trees and relative divergence dates in its input. A divergence time bound for species a and b is either an upper or lower bound (or both) on the number of years ago a and b diverged. To include this information as well as the other information provided by the other inputs, we use a dated phylogenetic tree. Such a tree is similar to that of a ranked phylogenetic tree in that values are assigned to the interior vertices except that these values now represent the number of years ago the corresponding speciation events occurred. Compatibility for this extended input is defined in the obvious way and RankedTree can be modified to solve this compatibility problem also.
6 4 Supertree Algorithms for Nested Taxa For supertree algorithms that take as their input collections of rooted phylogenetic trees and only return rooted trees that are leaf-labelled, it is implicit that, as a whole, the leaves of the trees in the input collection represent non-nested taxa. Thus, for example, Rattus rattus and mammal cannot be represented by two distinct leaves in such a collection as the former is nested inside the latter. This is somewhat limiting in the choice of trees for our input collection. In this section, we describe two supertree algorithms for combining rooted trees in which all of the leaves as well as some of the interior vertices are labelled. These trees are called rooted semi-labelled trees and the interior labels of such trees represent taxa at a level higher than that of their descendants. Two semi-labelled trees are shown in Fig. 3. g h g a b f c d e T a b c d e T Fig. 3. Two semi-labelled trees. The two algorithms for combining collections of rooted semi-labelled trees are called Semi-LabelledBuild and AncestralBuild. Both algorithms allow a leaf of one of the input trees to represent a taxa that is represented by an interior label of another tree. The motivation for both algorithms came from a problem posed by Page (2004). 4.1 Semi-LabelledBuild We say that a rooted semi-labelled tree T perfectly displays a rooted semi-labelled tree T if the label set X of T is a subset of the label set of T and, up to suppressing degree-two vertices, T is the minimal rooted subtree of T that connects the labels in X. Intuitively, T perfectly displays T if T preserves all of the ancestral relationships described by T exactly. In particular, T preserves all of the most recent common ancestor relationships described by T. A collection P of rooted semi-labelled trees is perfectly compatible if there is a rooted semilabelled tree T that perfectly displays each of the trees in P.
7 For a collection P of rooted semi-labelled trees, Semi-LabelledBuild decides whether or not P is perfectly compatible. Moreover, if P is perfectly compatible, then Semi-LabelledBuild returns a rooted semi-labelled tree that perfectly displays each of the trees in P. Figure 4 shows an application of Semi- LabelledBuild. The input consists of the two rooted semi-labelled trees shown in Fig. 4(a) and (b). Both input trees describe the evolution of spiders and were obtained from study S1xx9c14c42c30 in TreeBASE. There are taxa common to both trees and it is of particular interest to note that the taxon Araneoclada labels a leaf in the tree in (a), but an interior vertex in the tree in (b). The rooted semi-labelled tree resulting from applying Semi-LabelledBuild to the two input trees is shown in Fig. 4(c). Gradungulidae Scytodoidea (a) Neocribellatae Austrochilidae (b) Filistatidae Amaurobioidea Araneomorphae Araneoclada Lycosoidea Eresidae Opisthothelae Hypochilidae Araneoclada Oecobiidae Deinopidae Araneae Mygalomorphae Neocribellatae Orbiculariae Uloboridae Araneoidea Arachnida Liphistiomorphae Araneomorphae Dictynoidea Austrochilidae Amblypygi Paleocribellatae (c) Araneoclada Orbiculariae Neocribellatae Araneomorphae Opisthothelae Araneae Arachnida Scytodoidea Filistatidae Amaurobioidea Lycosoidea Eresidae Oecobiidae Uloboridae Deinopidae Araneoidea Dictynoidea Austrochilidae Gradungulidae Paleocribellatae Hypochilidae Mygalomorphae Liphistiomorphae Amblypygi Fig. 4. An application of Semi-LabelledBuild. 4.2 AncestralBuild The criteria of perfectly displays is very strong as a collection P of rooted semilabelled trees is perfectly compatible precisely if there is a rooted semi-labelled
8 tree T that preserves all of the most recent common ancestor relationships described by the collection. Thus Semi-LabelledBuild does not allow for the resolution of polytomies. The compatibility criteria and the associated algorithm we describe next relaxes this criteria and allows for the resolution of polytomies. A rooted semi-labelled tree T ancestrally displays a rooted semi-labelled tree T if the following properties hold: (i) the label set X of T is a subset of the label set of T ; (ii) up to suppressing degree-two vertices, T is a refinement of the minimal rooted subtree of T that connects the labels in X ; and (iii) for all labels in X, (a) if a is a proper ancestor of b in T, then a is a proper ancestor of b in T, and (b) if a is neither an ancestor nor a descendant of b in T, then a is neither an ancestor nor a descendant of b in T. To illustrate ancestrally displays and compare it with perfectly displays, in Fig. 3, T ancestrally displays T, but T does not perfectly display T. One can think of ancestrally displays as preserving all of the ancestor-descendant relationships. However, observe that it does not preserve the most recent common ancestor relationships. For example, if the most recent common ancestor of a and b is c in T, then, although c is an ancestor of both a and b in T, and neither a nor b is an ancestor of each other in T, c need not be the most recent common ancestor of a and b in T. A collection P of rooted semi-labelled trees is ancestrally compatible if there is a rooted semi-labelled tree T that ancestrally displays each of the trees in P. The algorithm AncestralBuild determines the ancestral compatibility of a collection of rooted semi-labelled trees, in which case, it outputs a rooted semi-labelled tree that ancestrally displays each of the trees in this collection. 5 Conclusion Supertree methods have attracted much interest recently, particularly in the light of well-funded Tree of Life initiatives, and studies that have combined large numbers of trees to construct phylogenies on hundreds, or even thousands of species. This has led to some vigorous argument both for and against the use of supertree (versus supermatrix) approaches for phylogeny reconstruction, as well as the emergence of some new techniques as alternatives to the standard MRP (matrix recoding with parsimony) approach. In this paper, we have described some further new methods, that allow for additional types of information to
9 be incorporated information that would be difficult to include directly using a traditional MRP analysis. It is important to note that all of the algorithms described in this paper are all-or-nothing algorithms. Each algorithm either returns a supertree with certain desirable properties relative to the input or returns a statement indicating that there is no such supertree. In practice, this limits their use. However, such algorithms are important first steps in developing supertree algorithms that always return a supertree and whose input includes information that goes beyond leaf-labelled phylogenetic trees. Indeed, for each of the algorithms described in this paper, we are currently developing MinCutSupertree type algorithms (Page, 2002; Semple and Steel, 2000) that overcomes this limitation. Lastly, if a supertree is returned by one of the algorithms described in this paper, two natural questions arise: (i) how many such supertrees are there and (ii) what common information is carried by all of these supertrees? In the case of (i), if there are too many, one may want to refine the original data to reduce this number. However, it is an immediate consequence of the main result by Bordewich et al. (in press) that determining this number exactly is #P-complete for all of the algorithms described in this paper. For (ii), Daniel (2004) has investigated this problem in the case where the input consists of just rooted phylogenetic trees. An approach similar to that taken by Daniel could be used for the algorithms described in this paper. Acknowledgements. The second author was supported by the New Zealand Institute of Mathematics and its Applications funded programme Phylogenetic Genomics and the first and last authors were supported by the New Zealand Marsden Fund. References 1. Aho, A. V., Sagiv, Y., Szymanski, T. G., and Ullman, J. D. (191). Inferring a tree from lowest common ancestors with an application to the optimization of relational expressions, SIAM Journal on Computing, 10, Bininda-Emonds, O. R. P., ed. Phylogenetic supertrees: combining information to reveal the Tree of Life, in press. 3. Bordewich, M., Semple, C., and Talbot, J. Counting consistent phylogenetic trees is #P-complete, Advances in Applied Mathematics, in press. 4. Bryant, D., Semple, C., and Steel, M. Combining evolutionary trees with ancestral divergence dates. In O. Bininda-Emonds (ed.), Phylogenetic supertrees: combining information to reveal the Tree of Life, Computational Biology Series, Kluwer, in press. 5. Daniel P. (2004). Supertree Methods: Some New Approaches, MSc thesis, University of Canterbury.. Daniel, P. and Semple, C. Supertree algorithms for nested taxa. In O. Bininda- Emonds (ed.), Phylogenetic supertrees: combining information to reveal the Tree of Life, Computational Biology Series, Kluwer, in press.
10 . Janczewski, D. N., Modi, W. S., Stephens, J. C., and O Brien, S. J. (1995). Molecular Evolution of Mitochondrial 12S RNA and Cytochrome b Sequences in the Pantherine Lineage of Felidae, Molecular Biology and Evolution, 12, Page, R. D. M. (2002). Modified mincut supertrees. In R. Guigó and D. Gusfield (eds.), Proceedings of the Second International Workshop on Algorithms in Bioinformatics (WABI 2002), pp , Springer. 9. Page, R. D. M. Taxonomy, Supertrees, and the Tree of Life. In O. Bininda-Emonds (ed.), Phylogenetic supertrees: combining information to reveal the Tree of Life, Computational Biology Series, Kluwer, in press. 10. Semple, C. and Steel, M. (2000). A supertree method for rooted trees, Discrete Applied Mathematics, 105, Semple, C. and Steel, M. (2003). Phylogenetics, Oxford University Press. 12. Slattery, J. P., Johnson, W. E., Goldman, D., O Brien, S. J. (1994). Phylogenetic Reconstruction of South American Felids Defined by Protein Electrophoresis, Journal of Molecular Evolution, 39,
Charles Semple, Philip Daniel, Wim Hordijk, Roderic D M Page, and Mike Steel
 SUPERTREE ALGORITHMS FOR ANCESTRAL DIVERGENCE DATES AND NESTED TAXA Charles Semple, Philip Daniel, Wim Hordijk, Roderic D M Page, and Mike Steel Department of Mathematics and Statistics University of Canterbury
SUPERTREE ALGORITHMS FOR ANCESTRAL DIVERGENCE DATES AND NESTED TAXA Charles Semple, Philip Daniel, Wim Hordijk, Roderic D M Page, and Mike Steel Department of Mathematics and Statistics University of Canterbury
A 3-APPROXIMATION ALGORITHM FOR THE SUBTREE DISTANCE BETWEEN PHYLOGENIES. 1. Introduction
 A 3-APPROXIMATION ALGORITHM FOR THE SUBTREE DISTANCE BETWEEN PHYLOGENIES MAGNUS BORDEWICH 1, CATHERINE MCCARTIN 2, AND CHARLES SEMPLE 3 Abstract. In this paper, we give a (polynomial-time) 3-approximation
A 3-APPROXIMATION ALGORITHM FOR THE SUBTREE DISTANCE BETWEEN PHYLOGENIES MAGNUS BORDEWICH 1, CATHERINE MCCARTIN 2, AND CHARLES SEMPLE 3 Abstract. In this paper, we give a (polynomial-time) 3-approximation
NOTE ON THE HYBRIDIZATION NUMBER AND SUBTREE DISTANCE IN PHYLOGENETICS
 NOTE ON THE HYBRIDIZATION NUMBER AND SUBTREE DISTANCE IN PHYLOGENETICS PETER J. HUMPHRIES AND CHARLES SEMPLE Abstract. For two rooted phylogenetic trees T and T, the rooted subtree prune and regraft distance
NOTE ON THE HYBRIDIZATION NUMBER AND SUBTREE DISTANCE IN PHYLOGENETICS PETER J. HUMPHRIES AND CHARLES SEMPLE Abstract. For two rooted phylogenetic trees T and T, the rooted subtree prune and regraft distance
THE THREE-STATE PERFECT PHYLOGENY PROBLEM REDUCES TO 2-SAT
 COMMUNICATIONS IN INFORMATION AND SYSTEMS c 2009 International Press Vol. 9, No. 4, pp. 295-302, 2009 001 THE THREE-STATE PERFECT PHYLOGENY PROBLEM REDUCES TO 2-SAT DAN GUSFIELD AND YUFENG WU Abstract.
COMMUNICATIONS IN INFORMATION AND SYSTEMS c 2009 International Press Vol. 9, No. 4, pp. 295-302, 2009 001 THE THREE-STATE PERFECT PHYLOGENY PROBLEM REDUCES TO 2-SAT DAN GUSFIELD AND YUFENG WU Abstract.
RECOVERING A PHYLOGENETIC TREE USING PAIRWISE CLOSURE OPERATIONS
 RECOVERING A PHYLOGENETIC TREE USING PAIRWISE CLOSURE OPERATIONS KT Huber, V Moulton, C Semple, and M Steel Department of Mathematics and Statistics University of Canterbury Private Bag 4800 Christchurch,
RECOVERING A PHYLOGENETIC TREE USING PAIRWISE CLOSURE OPERATIONS KT Huber, V Moulton, C Semple, and M Steel Department of Mathematics and Statistics University of Canterbury Private Bag 4800 Christchurch,
Phylogenetic Networks, Trees, and Clusters
 Phylogenetic Networks, Trees, and Clusters Luay Nakhleh 1 and Li-San Wang 2 1 Department of Computer Science Rice University Houston, TX 77005, USA nakhleh@cs.rice.edu 2 Department of Biology University
Phylogenetic Networks, Trees, and Clusters Luay Nakhleh 1 and Li-San Wang 2 1 Department of Computer Science Rice University Houston, TX 77005, USA nakhleh@cs.rice.edu 2 Department of Biology University
A Phylogenetic Network Construction due to Constrained Recombination
 A Phylogenetic Network Construction due to Constrained Recombination Mohd. Abdul Hai Zahid Research Scholar Research Supervisors: Dr. R.C. Joshi Dr. Ankush Mittal Department of Electronics and Computer
A Phylogenetic Network Construction due to Constrained Recombination Mohd. Abdul Hai Zahid Research Scholar Research Supervisors: Dr. R.C. Joshi Dr. Ankush Mittal Department of Electronics and Computer
A CLUSTER REDUCTION FOR COMPUTING THE SUBTREE DISTANCE BETWEEN PHYLOGENIES
 A CLUSTER REDUCTION FOR COMPUTING THE SUBTREE DISTANCE BETWEEN PHYLOGENIES SIMONE LINZ AND CHARLES SEMPLE Abstract. Calculating the rooted subtree prune and regraft (rspr) distance between two rooted binary
A CLUSTER REDUCTION FOR COMPUTING THE SUBTREE DISTANCE BETWEEN PHYLOGENIES SIMONE LINZ AND CHARLES SEMPLE Abstract. Calculating the rooted subtree prune and regraft (rspr) distance between two rooted binary
NJMerge: A generic technique for scaling phylogeny estimation methods and its application to species trees
 NJMerge: A generic technique for scaling phylogeny estimation methods and its application to species trees Erin Molloy and Tandy Warnow {emolloy2, warnow}@illinois.edu University of Illinois at Urbana
NJMerge: A generic technique for scaling phylogeny estimation methods and its application to species trees Erin Molloy and Tandy Warnow {emolloy2, warnow}@illinois.edu University of Illinois at Urbana
Chapter 26: Phylogeny and the Tree of Life Phylogenies Show Evolutionary Relationships
 Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
RECOVERING NORMAL NETWORKS FROM SHORTEST INTER-TAXA DISTANCE INFORMATION
 RECOVERING NORMAL NETWORKS FROM SHORTEST INTER-TAXA DISTANCE INFORMATION MAGNUS BORDEWICH, KATHARINA T. HUBER, VINCENT MOULTON, AND CHARLES SEMPLE Abstract. Phylogenetic networks are a type of leaf-labelled,
RECOVERING NORMAL NETWORKS FROM SHORTEST INTER-TAXA DISTANCE INFORMATION MAGNUS BORDEWICH, KATHARINA T. HUBER, VINCENT MOULTON, AND CHARLES SEMPLE Abstract. Phylogenetic networks are a type of leaf-labelled,
arxiv: v1 [q-bio.pe] 1 Jun 2014
![arxiv: v1 [q-bio.pe] 1 Jun 2014 arxiv: v1 [q-bio.pe] 1 Jun 2014](/thumbs/90/101381538.jpg) THE MOST PARSIMONIOUS TREE FOR RANDOM DATA MAREIKE FISCHER, MICHELLE GALLA, LINA HERBST AND MIKE STEEL arxiv:46.27v [q-bio.pe] Jun 24 Abstract. Applying a method to reconstruct a phylogenetic tree from
THE MOST PARSIMONIOUS TREE FOR RANDOM DATA MAREIKE FISCHER, MICHELLE GALLA, LINA HERBST AND MIKE STEEL arxiv:46.27v [q-bio.pe] Jun 24 Abstract. Applying a method to reconstruct a phylogenetic tree from
arxiv: v1 [q-bio.pe] 6 Jun 2013
![arxiv: v1 [q-bio.pe] 6 Jun 2013 arxiv: v1 [q-bio.pe] 6 Jun 2013](/thumbs/79/79187519.jpg) Hide and see: placing and finding an optimal tree for thousands of homoplasy-rich sequences Dietrich Radel 1, Andreas Sand 2,3, and Mie Steel 1, 1 Biomathematics Research Centre, University of Canterbury,
Hide and see: placing and finding an optimal tree for thousands of homoplasy-rich sequences Dietrich Radel 1, Andreas Sand 2,3, and Mie Steel 1, 1 Biomathematics Research Centre, University of Canterbury,
C.DARWIN ( )
 C.DARWIN (1809-1882) LAMARCK Each evolutionary lineage has evolved, transforming itself, from a ancestor appeared by spontaneous generation DARWIN All organisms are historically interconnected. Their relationships
C.DARWIN (1809-1882) LAMARCK Each evolutionary lineage has evolved, transforming itself, from a ancestor appeared by spontaneous generation DARWIN All organisms are historically interconnected. Their relationships
BIOINFORMATICS GABRIEL VALIENTE ALGORITHMS, BIOINFORMATICS, COMPLEXITY AND FORMAL METHODS RESEARCH GROUP, TECHNICAL UNIVERSITY OF CATALONIA
 BIOINFORMATICS GABRIEL VALIENTE ALGORITHMS, BIOINFORMATICS, COMPLEXITY AND FORMAL METHODS RESEARCH GROUP, TECHNICAL UNIVERSITY OF CATALONIA 2005 2006 Gabriel Valiente (ALBCOM) Bioinformatics 2005 2006
BIOINFORMATICS GABRIEL VALIENTE ALGORITHMS, BIOINFORMATICS, COMPLEXITY AND FORMAL METHODS RESEARCH GROUP, TECHNICAL UNIVERSITY OF CATALONIA 2005 2006 Gabriel Valiente (ALBCOM) Bioinformatics 2005 2006
Properties of normal phylogenetic networks
 Properties of normal phylogenetic networks Stephen J. Willson Department of Mathematics Iowa State University Ames, IA 50011 USA swillson@iastate.edu August 13, 2009 Abstract. A phylogenetic network is
Properties of normal phylogenetic networks Stephen J. Willson Department of Mathematics Iowa State University Ames, IA 50011 USA swillson@iastate.edu August 13, 2009 Abstract. A phylogenetic network is
arxiv: v1 [cs.cc] 9 Oct 2014
![arxiv: v1 [cs.cc] 9 Oct 2014 arxiv: v1 [cs.cc] 9 Oct 2014](/thumbs/78/77800744.jpg) Satisfying ternary permutation constraints by multiple linear orders or phylogenetic trees Leo van Iersel, Steven Kelk, Nela Lekić, Simone Linz May 7, 08 arxiv:40.7v [cs.cc] 9 Oct 04 Abstract A ternary
Satisfying ternary permutation constraints by multiple linear orders or phylogenetic trees Leo van Iersel, Steven Kelk, Nela Lekić, Simone Linz May 7, 08 arxiv:40.7v [cs.cc] 9 Oct 04 Abstract A ternary
Macroevolution Part I: Phylogenies
 Macroevolution Part I: Phylogenies Taxonomy Classification originated with Carolus Linnaeus in the 18 th century. Based on structural (outward and inward) similarities Hierarchal scheme, the largest most
Macroevolution Part I: Phylogenies Taxonomy Classification originated with Carolus Linnaeus in the 18 th century. Based on structural (outward and inward) similarities Hierarchal scheme, the largest most
8/23/2014. Phylogeny and the Tree of Life
 Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley
 Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Biological Networks: Comparison, Conservation, and Evolution via Relative Description Length By: Tamir Tuller & Benny Chor
 Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
PHYLOGENY AND SYSTEMATICS
 AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
arxiv: v1 [q-bio.pe] 16 Aug 2007
![arxiv: v1 [q-bio.pe] 16 Aug 2007 arxiv: v1 [q-bio.pe] 16 Aug 2007](/thumbs/95/125806968.jpg) MAXIMUM LIKELIHOOD SUPERTREES arxiv:0708.2124v1 [q-bio.pe] 16 Aug 2007 MIKE STEEL AND ALLEN RODRIGO Abstract. We analyse a maximum-likelihood approach for combining phylogenetic trees into a larger supertree.
MAXIMUM LIKELIHOOD SUPERTREES arxiv:0708.2124v1 [q-bio.pe] 16 Aug 2007 MIKE STEEL AND ALLEN RODRIGO Abstract. We analyse a maximum-likelihood approach for combining phylogenetic trees into a larger supertree.
Anatomy of a tree. clade is group of organisms with a shared ancestor. a monophyletic group shares a single common ancestor = tapirs-rhinos-horses
 Anatomy of a tree outgroup: an early branching relative of the interest groups sister taxa: taxa derived from the same recent ancestor polytomy: >2 taxa emerge from a node Anatomy of a tree clade is group
Anatomy of a tree outgroup: an early branching relative of the interest groups sister taxa: taxa derived from the same recent ancestor polytomy: >2 taxa emerge from a node Anatomy of a tree clade is group
DISTRIBUTIONS OF CHERRIES FOR TWO MODELS OF TREES
 DISTRIBUTIONS OF CHERRIES FOR TWO MODELS OF TREES ANDY McKENZIE and MIKE STEEL Biomathematics Research Centre University of Canterbury Private Bag 4800 Christchurch, New Zealand No. 177 May, 1999 Distributions
DISTRIBUTIONS OF CHERRIES FOR TWO MODELS OF TREES ANDY McKENZIE and MIKE STEEL Biomathematics Research Centre University of Canterbury Private Bag 4800 Christchurch, New Zealand No. 177 May, 1999 Distributions
Outline. Classification of Living Things
 Outline Classification of Living Things Chapter 20 Mader: Biology 8th Ed. Taxonomy Binomial System Species Identification Classification Categories Phylogenetic Trees Tracing Phylogeny Cladistic Systematics
Outline Classification of Living Things Chapter 20 Mader: Biology 8th Ed. Taxonomy Binomial System Species Identification Classification Categories Phylogenetic Trees Tracing Phylogeny Cladistic Systematics
BINF6201/8201. Molecular phylogenetic methods
 BINF60/80 Molecular phylogenetic methods 0-7-06 Phylogenetics Ø According to the evolutionary theory, all life forms on this planet are related to one another by descent. Ø Traditionally, phylogenetics
BINF60/80 Molecular phylogenetic methods 0-7-06 Phylogenetics Ø According to the evolutionary theory, all life forms on this planet are related to one another by descent. Ø Traditionally, phylogenetics
A (short) introduction to phylogenetics
 A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
Phylogenetic Tree Reconstruction
 I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
UoN, CAS, DBSC BIOL102 lecture notes by: Dr. Mustafa A. Mansi. The Phylogenetic Systematics (Phylogeny and Systematics)
 - Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
- Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
Inferring a level-1 phylogenetic network from a dense set of rooted triplets
 Theoretical Computer Science 363 (2006) 60 68 www.elsevier.com/locate/tcs Inferring a level-1 phylogenetic network from a dense set of rooted triplets Jesper Jansson a,, Wing-Kin Sung a,b, a School of
Theoretical Computer Science 363 (2006) 60 68 www.elsevier.com/locate/tcs Inferring a level-1 phylogenetic network from a dense set of rooted triplets Jesper Jansson a,, Wing-Kin Sung a,b, a School of
Reconstructing the history of lineages
 Reconstructing the history of lineages Class outline Systematics Phylogenetic systematics Phylogenetic trees and maps Class outline Definitions Systematics Phylogenetic systematics/cladistics Systematics
Reconstructing the history of lineages Class outline Systematics Phylogenetic systematics Phylogenetic trees and maps Class outline Definitions Systematics Phylogenetic systematics/cladistics Systematics
A 3-approximation algorithm for the subtree distance between phylogenies
 JID:JDA AID:228 /FLA [m3sc+; v 1.87; Prn:5/02/2008; 10:04] P.1 (1-14) Journal of Discrete Algorithms ( ) www.elsevier.com/locate/jda A 3-approximation algorithm for the subtree distance between phylogenies
JID:JDA AID:228 /FLA [m3sc+; v 1.87; Prn:5/02/2008; 10:04] P.1 (1-14) Journal of Discrete Algorithms ( ) www.elsevier.com/locate/jda A 3-approximation algorithm for the subtree distance between phylogenies
Non-binary Tree Reconciliation. Louxin Zhang Department of Mathematics National University of Singapore
 Non-binary Tree Reconciliation Louxin Zhang Department of Mathematics National University of Singapore matzlx@nus.edu.sg Introduction: Gene Duplication Inference Consider a duplication gene family G Species
Non-binary Tree Reconciliation Louxin Zhang Department of Mathematics National University of Singapore matzlx@nus.edu.sg Introduction: Gene Duplication Inference Consider a duplication gene family G Species
Phylogeny 9/8/2014. Evolutionary Relationships. Data Supporting Phylogeny. Chapter 26
 Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Phylogenetic Trees. What They Are Why We Do It & How To Do It. Presented by Amy Harris Dr Brad Morantz
 Phylogenetic Trees What They Are Why We Do It & How To Do It Presented by Amy Harris Dr Brad Morantz Overview What is a phylogenetic tree Why do we do it How do we do it Methods and programs Parallels
Phylogenetic Trees What They Are Why We Do It & How To Do It Presented by Amy Harris Dr Brad Morantz Overview What is a phylogenetic tree Why do we do it How do we do it Methods and programs Parallels
Parsimony via Consensus
 Syst. Biol. 57(2):251 256, 2008 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150802040597 Parsimony via Consensus TREVOR C. BRUEN 1 AND DAVID
Syst. Biol. 57(2):251 256, 2008 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150802040597 Parsimony via Consensus TREVOR C. BRUEN 1 AND DAVID
What is Phylogenetics
 What is Phylogenetics Phylogenetics is the area of research concerned with finding the genetic connections and relationships between species. The basic idea is to compare specific characters (features)
What is Phylogenetics Phylogenetics is the area of research concerned with finding the genetic connections and relationships between species. The basic idea is to compare specific characters (features)
Chapter 26 Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Chapter focus Shifting from the process of how evolution works to the pattern evolution produces over time. Phylogeny Phylon = tribe, geny = genesis or origin
Chapter 26 Phylogeny and the Tree of Life Chapter focus Shifting from the process of how evolution works to the pattern evolution produces over time. Phylogeny Phylon = tribe, geny = genesis or origin
Regular networks are determined by their trees
 Regular networks are determined by their trees Stephen J. Willson Department of Mathematics Iowa State University Ames, IA 50011 USA swillson@iastate.edu February 17, 2009 Abstract. A rooted acyclic digraph
Regular networks are determined by their trees Stephen J. Willson Department of Mathematics Iowa State University Ames, IA 50011 USA swillson@iastate.edu February 17, 2009 Abstract. A rooted acyclic digraph
Lecture 11 Friday, October 21, 2011
 Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
Copyright (c) 2008 Daniel Huson. Permission is granted to copy, distribute and/or modify this document under the terms of the GNU Free Documentation
 Copyright (c) 2008 Daniel Huson. Permission is granted to copy, distribute and/or modify this document under the terms of the GNU Free Documentation License, Version 1.2 or any later version published
Copyright (c) 2008 Daniel Huson. Permission is granted to copy, distribute and/or modify this document under the terms of the GNU Free Documentation License, Version 1.2 or any later version published
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Minimal Phylogenetic Supertrees and Local Consensus Trees
 Minimal Phylogenetic Supertrees and Local Consensus Trees Jesper Jansson 1 and Wing-Kin Sung 1 Laboratory of Mathematical Bioinformatics, Institute for Chemical Research, Kyoto University, Kyoto, Japan
Minimal Phylogenetic Supertrees and Local Consensus Trees Jesper Jansson 1 and Wing-Kin Sung 1 Laboratory of Mathematical Bioinformatics, Institute for Chemical Research, Kyoto University, Kyoto, Japan
BIOL 1010 Introduction to Biology: The Evolution and Diversity of Life. Spring 2011 Sections A & B
 BIOL 1010 Introduction to Biology: The Evolution and Diversity of Life. Spring 2011 Sections A & B Steve Thompson: stthompson@valdosta.edu http://www.bioinfo4u.net 1 ʻTree of Life,ʼ ʻprimitive,ʼ ʻprogressʼ
BIOL 1010 Introduction to Biology: The Evolution and Diversity of Life. Spring 2011 Sections A & B Steve Thompson: stthompson@valdosta.edu http://www.bioinfo4u.net 1 ʻTree of Life,ʼ ʻprimitive,ʼ ʻprogressʼ
What is the purpose of the Classifying System? To allow the accurate identification of a particular organism
 What is the purpose of the Classifying System? To allow the accurate identification of a particular organism Taxonomy The practice of classifying organisms -Taxonomy was founded nearly 300 years ago by
What is the purpose of the Classifying System? To allow the accurate identification of a particular organism Taxonomy The practice of classifying organisms -Taxonomy was founded nearly 300 years ago by
C3020 Molecular Evolution. Exercises #3: Phylogenetics
 C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
Comparative Genomics II
 Comparative Genomics II Advances in Bioinformatics and Genomics GEN 240B Jason Stajich May 19 Comparative Genomics II Slide 1/31 Outline Introduction Gene Families Pairwise Methods Phylogenetic Methods
Comparative Genomics II Advances in Bioinformatics and Genomics GEN 240B Jason Stajich May 19 Comparative Genomics II Slide 1/31 Outline Introduction Gene Families Pairwise Methods Phylogenetic Methods
AP Biology. Cladistics
 Cladistics Kingdom Summary Review slide Review slide Classification Old 5 Kingdom system Eukaryote Monera, Protists, Plants, Fungi, Animals New 3 Domain system reflects a greater understanding of evolution
Cladistics Kingdom Summary Review slide Review slide Classification Old 5 Kingdom system Eukaryote Monera, Protists, Plants, Fungi, Animals New 3 Domain system reflects a greater understanding of evolution
CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny
 CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny To trace phylogeny or the evolutionary history of life, biologists use evidence from paleontology, molecular data, comparative
CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny To trace phylogeny or the evolutionary history of life, biologists use evidence from paleontology, molecular data, comparative
X X (2) X Pr(X = x θ) (3)
 Notes for 848 lecture 6: A ML basis for compatibility and parsimony Notation θ Θ (1) Θ is the space of all possible trees (and model parameters) θ is a point in the parameter space = a particular tree
Notes for 848 lecture 6: A ML basis for compatibility and parsimony Notation θ Θ (1) Θ is the space of all possible trees (and model parameters) θ is a point in the parameter space = a particular tree
Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 26 Phylogeny and the Tree of Life PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Phylogenetics. Applications of phylogenetics. Unrooted networks vs. rooted trees. Outline
 Phylogenetics Todd Vision iology 522 March 26, 2007 pplications of phylogenetics Studying organismal or biogeographic history Systematics ating events in the fossil record onservation biology Studying
Phylogenetics Todd Vision iology 522 March 26, 2007 pplications of phylogenetics Studying organismal or biogeographic history Systematics ating events in the fossil record onservation biology Studying
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics
 POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
Cherry picking: a characterization of the temporal hybridization number for a set of phylogenies
 Bulletin of Mathematical Biology manuscript No. (will be inserted by the editor) Cherry picking: a characterization of the temporal hybridization number for a set of phylogenies Peter J. Humphries Simone
Bulletin of Mathematical Biology manuscript No. (will be inserted by the editor) Cherry picking: a characterization of the temporal hybridization number for a set of phylogenies Peter J. Humphries Simone
Determining a Binary Matroid from its Small Circuits
 Determining a Binary Matroid from its Small Circuits James Oxley Department of Mathematics Louisiana State University Louisiana, USA oxley@math.lsu.edu Charles Semple School of Mathematics and Statistics
Determining a Binary Matroid from its Small Circuits James Oxley Department of Mathematics Louisiana State University Louisiana, USA oxley@math.lsu.edu Charles Semple School of Mathematics and Statistics
Beyond Galled Trees Decomposition and Computation of Galled Networks
 Beyond Galled Trees Decomposition and Computation of Galled Networks Daniel H. Huson & Tobias H.Kloepper RECOMB 2007 1 Copyright (c) 2008 Daniel Huson. Permission is granted to copy, distribute and/or
Beyond Galled Trees Decomposition and Computation of Galled Networks Daniel H. Huson & Tobias H.Kloepper RECOMB 2007 1 Copyright (c) 2008 Daniel Huson. Permission is granted to copy, distribute and/or
Chapter 26 Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Evolutionary Tree Analysis. Overview
 CSI/BINF 5330 Evolutionary Tree Analysis Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Backgrounds Distance-Based Evolutionary Tree Reconstruction Character-Based
CSI/BINF 5330 Evolutionary Tree Analysis Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Backgrounds Distance-Based Evolutionary Tree Reconstruction Character-Based
Phylogeny and systematics. Why are these disciplines important in evolutionary biology and how are they related to each other?
 Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Probabilities of Evolutionary Trees under a Rate-Varying Model of Speciation
 Probabilities of Evolutionary Trees under a Rate-Varying Model of Speciation Mike Steel Biomathematics Research Centre University of Canterbury, Private Bag 4800 Christchurch, New Zealand No. 67 December,
Probabilities of Evolutionary Trees under a Rate-Varying Model of Speciation Mike Steel Biomathematics Research Centre University of Canterbury, Private Bag 4800 Christchurch, New Zealand No. 67 December,
Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 26 Phylogeny and the Tree of Life PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Algorithmic aspects of tree amalgamation
 June 12, 2000 Algorithmic aspects of tree amalgamation Sebastian Böcker GK Strukturbildungsprozesse, FSP Mathematisierung, Universität Bielefeld, PF 100 131, 33501 Bielefeld, Germany boecker@mathematik.uni-bielefeld.de
June 12, 2000 Algorithmic aspects of tree amalgamation Sebastian Böcker GK Strukturbildungsprozesse, FSP Mathematisierung, Universität Bielefeld, PF 100 131, 33501 Bielefeld, Germany boecker@mathematik.uni-bielefeld.de
Phylogeny: traditional and Bayesian approaches
 Phylogeny: traditional and Bayesian approaches 5-Feb-2014 DEKM book Notes from Dr. B. John Holder and Lewis, Nature Reviews Genetics 4, 275-284, 2003 1 Phylogeny A graph depicting the ancestor-descendent
Phylogeny: traditional and Bayesian approaches 5-Feb-2014 DEKM book Notes from Dr. B. John Holder and Lewis, Nature Reviews Genetics 4, 275-284, 2003 1 Phylogeny A graph depicting the ancestor-descendent
UNICYCLIC NETWORKS: COMPATIBILITY AND ENUMERATION
 UNICYCLIC NETWORKS: COMPATIBILITY AND ENUMERATION CHARLES SEMPLE AND MIKE STEEL Abstract. Graphs obtained from a binary leaf labelled ( phylogenetic ) tree by adding an edge so as to introduce a cycle
UNICYCLIC NETWORKS: COMPATIBILITY AND ENUMERATION CHARLES SEMPLE AND MIKE STEEL Abstract. Graphs obtained from a binary leaf labelled ( phylogenetic ) tree by adding an edge so as to introduce a cycle
Using phylogenetics to estimate species divergence times... Basics and basic issues for Bayesian inference of divergence times (plus some digression)
 Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
arxiv: v1 [cs.ds] 1 Nov 2018
![arxiv: v1 [cs.ds] 1 Nov 2018 arxiv: v1 [cs.ds] 1 Nov 2018](/thumbs/90/101592235.jpg) An O(nlogn) time Algorithm for computing the Path-length Distance between Trees arxiv:1811.00619v1 [cs.ds] 1 Nov 2018 David Bryant Celine Scornavacca November 5, 2018 Abstract Tree comparison metrics have
An O(nlogn) time Algorithm for computing the Path-length Distance between Trees arxiv:1811.00619v1 [cs.ds] 1 Nov 2018 David Bryant Celine Scornavacca November 5, 2018 Abstract Tree comparison metrics have
Lecture V Phylogeny and Systematics Dr. Kopeny
 Delivered 1/30 and 2/1 Lecture V Phylogeny and Systematics Dr. Kopeny Lecture V How to Determine Evolutionary Relationships: Concepts in Phylogeny and Systematics Textbook Reading: pp 425-433, 435-437
Delivered 1/30 and 2/1 Lecture V Phylogeny and Systematics Dr. Kopeny Lecture V How to Determine Evolutionary Relationships: Concepts in Phylogeny and Systematics Textbook Reading: pp 425-433, 435-437
Axiomatic Opportunities and Obstacles for Inferring a Species Tree from Gene Trees
 Syst. Biol. 63(5):772 778, 2014 The Author(s) 2014. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Syst. Biol. 63(5):772 778, 2014 The Author(s) 2014. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
AN ALGORITHM FOR CONSTRUCTING A k-tree FOR A k-connected MATROID
 AN ALGORITHM FOR CONSTRUCTING A k-tree FOR A k-connected MATROID NICK BRETTELL AND CHARLES SEMPLE Dedicated to James Oxley on the occasion of his 60th birthday Abstract. For a k-connected matroid M, Clark
AN ALGORITHM FOR CONSTRUCTING A k-tree FOR A k-connected MATROID NICK BRETTELL AND CHARLES SEMPLE Dedicated to James Oxley on the occasion of his 60th birthday Abstract. For a k-connected matroid M, Clark
Non-independence in Statistical Tests for Discrete Cross-species Data
 J. theor. Biol. (1997) 188, 507514 Non-independence in Statistical Tests for Discrete Cross-species Data ALAN GRAFEN* AND MARK RIDLEY * St. John s College, Oxford OX1 3JP, and the Department of Zoology,
J. theor. Biol. (1997) 188, 507514 Non-independence in Statistical Tests for Discrete Cross-species Data ALAN GRAFEN* AND MARK RIDLEY * St. John s College, Oxford OX1 3JP, and the Department of Zoology,
(Stevens 1991) 1. morphological characters should be assumed to be quantitative unless demonstrated otherwise
 Bot 421/521 PHYLOGENETIC ANALYSIS I. Origins A. Hennig 1950 (German edition) Phylogenetic Systematics 1966 B. Zimmerman (Germany, 1930 s) C. Wagner (Michigan, 1920-2000) II. Characters and character states
Bot 421/521 PHYLOGENETIC ANALYSIS I. Origins A. Hennig 1950 (German edition) Phylogenetic Systematics 1966 B. Zimmerman (Germany, 1930 s) C. Wagner (Michigan, 1920-2000) II. Characters and character states
Consistency Index (CI)
 Consistency Index (CI) minimum number of changes divided by the number required on the tree. CI=1 if there is no homoplasy negatively correlated with the number of species sampled Retention Index (RI)
Consistency Index (CI) minimum number of changes divided by the number required on the tree. CI=1 if there is no homoplasy negatively correlated with the number of species sampled Retention Index (RI)
Let S be a set of n species. A phylogeny is a rooted tree with n leaves, each of which is uniquely
 JOURNAL OF COMPUTATIONAL BIOLOGY Volume 8, Number 1, 2001 Mary Ann Liebert, Inc. Pp. 69 78 Perfect Phylogenetic Networks with Recombination LUSHENG WANG, 1 KAIZHONG ZHANG, 2 and LOUXIN ZHANG 3 ABSTRACT
JOURNAL OF COMPUTATIONAL BIOLOGY Volume 8, Number 1, 2001 Mary Ann Liebert, Inc. Pp. 69 78 Perfect Phylogenetic Networks with Recombination LUSHENG WANG, 1 KAIZHONG ZHANG, 2 and LOUXIN ZHANG 3 ABSTRACT
CONSTRUCTING A 3-TREE FOR A 3-CONNECTED MATROID. 1. Introduction
 CONSTRUCTING A 3-TREE FOR A 3-CONNECTED MATROID JAMES OXLEY AND CHARLES SEMPLE For our friend Geoff Whittle with thanks for many years of enjoyable collaboration Abstract. In an earlier paper with Whittle,
CONSTRUCTING A 3-TREE FOR A 3-CONNECTED MATROID JAMES OXLEY AND CHARLES SEMPLE For our friend Geoff Whittle with thanks for many years of enjoyable collaboration Abstract. In an earlier paper with Whittle,
Biology 211 (2) Week 1 KEY!
 Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
arxiv: v5 [q-bio.pe] 24 Oct 2016
![arxiv: v5 [q-bio.pe] 24 Oct 2016 arxiv: v5 [q-bio.pe] 24 Oct 2016](/thumbs/95/125807017.jpg) On the Quirks of Maximum Parsimony and Likelihood on Phylogenetic Networks Christopher Bryant a, Mareike Fischer b, Simone Linz c, Charles Semple d arxiv:1505.06898v5 [q-bio.pe] 24 Oct 2016 a Statistics
On the Quirks of Maximum Parsimony and Likelihood on Phylogenetic Networks Christopher Bryant a, Mareike Fischer b, Simone Linz c, Charles Semple d arxiv:1505.06898v5 [q-bio.pe] 24 Oct 2016 a Statistics
The Tree of Life. Phylogeny
 The Tree of Life Phylogeny Phylogenetics Phylogenetic trees illustrate the evolutionary relationships among groups of organisms, or among a family of related nucleic acid or protein sequences Each branch
The Tree of Life Phylogeny Phylogenetics Phylogenetic trees illustrate the evolutionary relationships among groups of organisms, or among a family of related nucleic acid or protein sequences Each branch
Chapter 19: Taxonomy, Systematics, and Phylogeny
 Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
A new algorithm to construct phylogenetic networks from trees
 A new algorithm to construct phylogenetic networks from trees J. Wang College of Computer Science, Inner Mongolia University, Hohhot, Inner Mongolia, China Corresponding author: J. Wang E-mail: wangjuanangle@hit.edu.cn
A new algorithm to construct phylogenetic networks from trees J. Wang College of Computer Science, Inner Mongolia University, Hohhot, Inner Mongolia, China Corresponding author: J. Wang E-mail: wangjuanangle@hit.edu.cn
CHAPTERS 24-25: Evidence for Evolution and Phylogeny
 CHAPTERS 24-25: Evidence for Evolution and Phylogeny 1. For each of the following, indicate how it is used as evidence of evolution by natural selection or shown as an evolutionary trend: a. Paleontology
CHAPTERS 24-25: Evidence for Evolution and Phylogeny 1. For each of the following, indicate how it is used as evidence of evolution by natural selection or shown as an evolutionary trend: a. Paleontology
Classification and Phylogeny
 Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization)
 STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization) Laura Salter Kubatko Departments of Statistics and Evolution, Ecology, and Organismal Biology The Ohio State University kubatko.2@osu.edu
STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization) Laura Salter Kubatko Departments of Statistics and Evolution, Ecology, and Organismal Biology The Ohio State University kubatko.2@osu.edu
1.1 The (rooted, binary-character) Perfect-Phylogeny Problem
 Contents 1 Trees First 3 1.1 Rooted Perfect-Phylogeny...................... 3 1.1.1 Alternative Definitions.................... 5 1.1.2 The Perfect-Phylogeny Problem and Solution....... 7 1.2 Alternate,
Contents 1 Trees First 3 1.1 Rooted Perfect-Phylogeny...................... 3 1.1.1 Alternative Definitions.................... 5 1.1.2 The Perfect-Phylogeny Problem and Solution....... 7 1.2 Alternate,
arxiv: v1 [q-bio.pe] 4 Sep 2013
![arxiv: v1 [q-bio.pe] 4 Sep 2013 arxiv: v1 [q-bio.pe] 4 Sep 2013](/thumbs/88/117441027.jpg) Version dated: September 5, 2013 Predicting ancestral states in a tree arxiv:1309.0926v1 [q-bio.pe] 4 Sep 2013 Predicting the ancestral character changes in a tree is typically easier than predicting the
Version dated: September 5, 2013 Predicting ancestral states in a tree arxiv:1309.0926v1 [q-bio.pe] 4 Sep 2013 Predicting the ancestral character changes in a tree is typically easier than predicting the
reconciling trees Stefanie Hartmann postdoc, Todd Vision s lab University of North Carolina the data
 reconciling trees Stefanie Hartmann postdoc, Todd Vision s lab University of North Carolina 1 the data alignments and phylogenies for ~27,000 gene families from 140 plant species www.phytome.org publicly
reconciling trees Stefanie Hartmann postdoc, Todd Vision s lab University of North Carolina 1 the data alignments and phylogenies for ~27,000 gene families from 140 plant species www.phytome.org publicly
Solving the Maximum Agreement Subtree and Maximum Comp. Tree problems on bounded degree trees. Sylvain Guillemot, François Nicolas.
 Solving the Maximum Agreement Subtree and Maximum Compatible Tree problems on bounded degree trees LIRMM, Montpellier France 4th July 2006 Introduction The Mast and Mct problems: given a set of evolutionary
Solving the Maximum Agreement Subtree and Maximum Compatible Tree problems on bounded degree trees LIRMM, Montpellier France 4th July 2006 Introduction The Mast and Mct problems: given a set of evolutionary
Classification and Phylogeny
 Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Gene Families part 2. Review: Gene Families /727 Lecture 8. Protein family. (Multi)gene family
 Review: Gene Families Gene Families part 2 03 327/727 Lecture 8 What is a Case study: ian globin genes Gene trees and how they differ from species trees Homology, orthology, and paralogy Last tuesday 1
Review: Gene Families Gene Families part 2 03 327/727 Lecture 8 What is a Case study: ian globin genes Gene trees and how they differ from species trees Homology, orthology, and paralogy Last tuesday 1
Reconstruction of certain phylogenetic networks from their tree-average distances
 Reconstruction of certain phylogenetic networks from their tree-average distances Stephen J. Willson Department of Mathematics Iowa State University Ames, IA 50011 USA swillson@iastate.edu October 10,
Reconstruction of certain phylogenetic networks from their tree-average distances Stephen J. Willson Department of Mathematics Iowa State University Ames, IA 50011 USA swillson@iastate.edu October 10,
Phylogenetic Trees. Phylogenetic Trees Five. Phylogeny: Inference Tool. Phylogeny Terminology. Picture of Last Quagga. Importance of Phylogeny 5.
 Five Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu v Distance Methods v Character Methods v Molecular Clock v UPGMA v Maximum Parsimony
Five Sami Khuri Department of Computer Science San José State University San José, California, USA sami.khuri@sjsu.edu v Distance Methods v Character Methods v Molecular Clock v UPGMA v Maximum Parsimony
On the Subnet Prune and Regraft distance
 On the Subnet Prune and Regraft distance Jonathan Klawitter and Simone Linz Department of Computer Science, University of Auckland, New Zealand jo. klawitter@ gmail. com, s. linz@ auckland. ac. nz arxiv:805.07839v
On the Subnet Prune and Regraft distance Jonathan Klawitter and Simone Linz Department of Computer Science, University of Auckland, New Zealand jo. klawitter@ gmail. com, s. linz@ auckland. ac. nz arxiv:805.07839v
Inferring Phylogenetic Trees. Distance Approaches. Representing distances. in rooted and unrooted trees. The distance approach to phylogenies
 Inferring Phylogenetic Trees Distance Approaches Representing distances in rooted and unrooted trees The distance approach to phylogenies given: an n n matrix M where M ij is the distance between taxa
Inferring Phylogenetic Trees Distance Approaches Representing distances in rooted and unrooted trees The distance approach to phylogenies given: an n n matrix M where M ij is the distance between taxa
SPR Distance Computation for Unrooted Trees
 ORIGINAL RESEARCH SPR Distance Computation for Unrooted Trees Glenn Hickey, Frank Dehne, Andrew Rau-Chaplin and Christian Blouin School of Computer Science, Carleton University, Ottawa, Canada KS 5B. School
ORIGINAL RESEARCH SPR Distance Computation for Unrooted Trees Glenn Hickey, Frank Dehne, Andrew Rau-Chaplin and Christian Blouin School of Computer Science, Carleton University, Ottawa, Canada KS 5B. School
Computational approaches for functional genomics
 Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Is the equal branch length model a parsimony model?
 Table 1: n approximation of the probability of data patterns on the tree shown in figure?? made by dropping terms that do not have the minimal exponent for p. Terms that were dropped are shown in red;
Table 1: n approximation of the probability of data patterns on the tree shown in figure?? made by dropping terms that do not have the minimal exponent for p. Terms that were dropped are shown in red;
CLASSIFICATION OF LIVING THINGS. Chapter 18
 CLASSIFICATION OF LIVING THINGS Chapter 18 How many species are there? About 1.8 million species have been given scientific names Nearly 2/3 of which are insects 99% of all known animal species are smaller
CLASSIFICATION OF LIVING THINGS Chapter 18 How many species are there? About 1.8 million species have been given scientific names Nearly 2/3 of which are insects 99% of all known animal species are smaller
Autotrophs capture the light energy from sunlight and convert it to chemical energy they use for food.
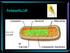 Prokaryotic Cell Eukaryotic Cell Autotrophs capture the light energy from sunlight and convert it to chemical energy they use for food. Heterotrophs must get energy by eating autotrophs or other heterotrophs.
Prokaryotic Cell Eukaryotic Cell Autotrophs capture the light energy from sunlight and convert it to chemical energy they use for food. Heterotrophs must get energy by eating autotrophs or other heterotrophs.
